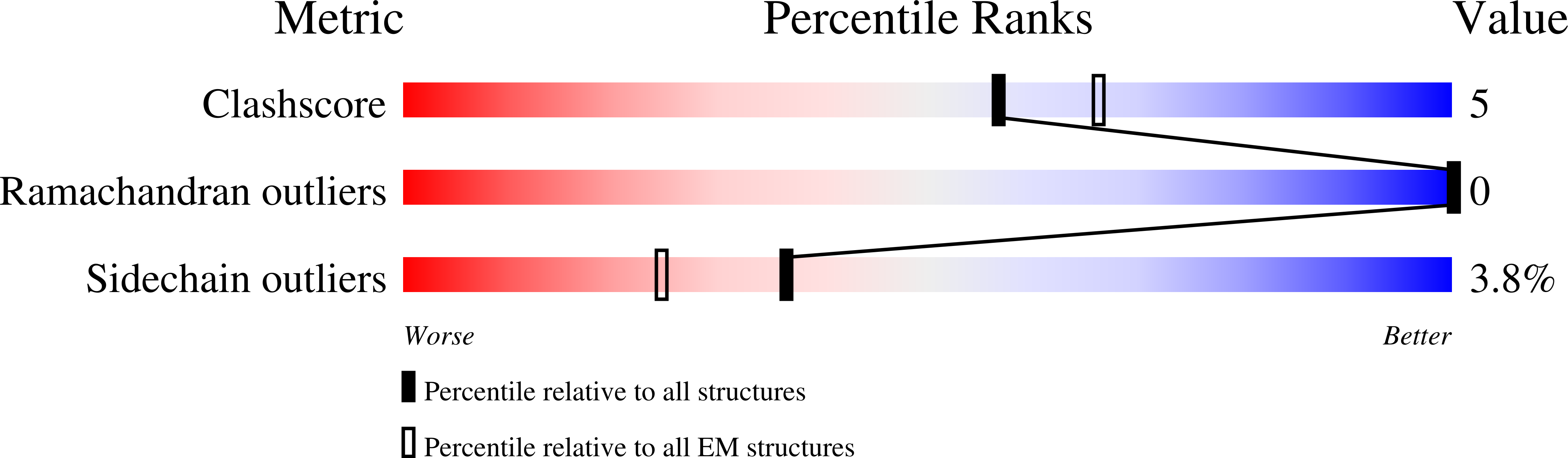Structural Basis for High-Affinity Trapping of the Na V 1.7 Channel in Its Resting State by Tarantula Toxin.
Wisedchaisri, G., Tonggu, L., Gamal El-Din, T.M., McCord, E., Zheng, N., Catterall, W.A.(2021) Mol Cell 81: 38-48.e4
- PubMed: 33232657
- DOI: https://doi.org/10.1016/j.molcel.2020.10.039
- Primary Citation of Related Structures:
7K48 - PubMed Abstract:
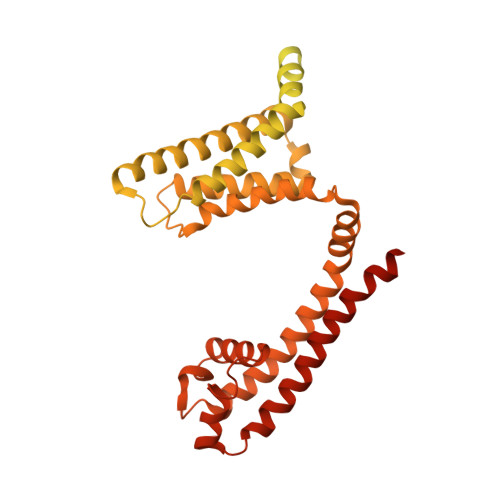
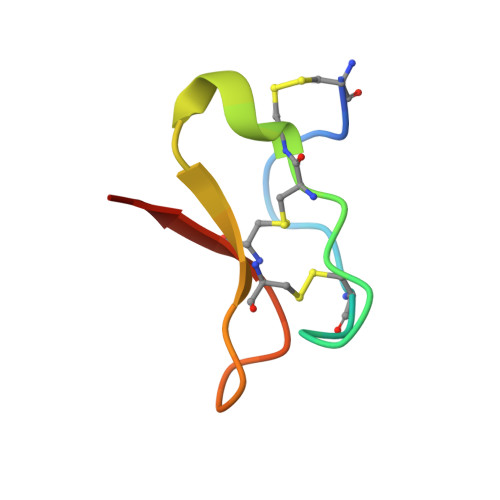
Voltage-gated sodium channels initiate electrical signals and are frequently targeted by deadly gating-modifier neurotoxins, including tarantula toxins, which trap the voltage sensor in its resting state. The structural basis for tarantula-toxin action remains elusive because of the difficulty of capturing the functionally relevant form of the toxin-channel complex. Here, we engineered the model sodium channel Na V Ab with voltage-shifting mutations and the toxin-binding site of human Na V 1.7, an attractive pain target. This mutant chimera enabled us to determine the cryoelectron microscopy (cryo-EM) structure of the channel functionally arrested by tarantula toxin. Our structure reveals a high-affinity resting-state-specific toxin-channel interaction between a key lysine residue that serves as a "stinger" and penetrates a triad of carboxyl groups in the S3-S4 linker of the voltage sensor. By unveiling this high-affinity binding mode, our studies establish a high-resolution channel-docking and resting-state locking mechanism for huwentoxin-IV and provide guidance for developing future resting-state-targeted analgesic drugs.
Organizational Affiliation:
Department of Pharmacology, University of Washington, Seattle, WA 98195, USA.