Structural insights into the potency and selectivity of covalent pan-FGFR inhibitors
Qu, L., Chen, X., Wei, H., Guo, M., Dai, S., Jiang, L., Li, J., Yue, S., Chen, Z., Chen, Y.(2022) Commun Chem 5: 5
Experimental Data Snapshot
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Fibroblast growth factor receptor 4 | 311 | Homo sapiens | Mutation(s): 1 Gene Names: FGFR4, JTK2, TKF EC: 2.7.10.1 | 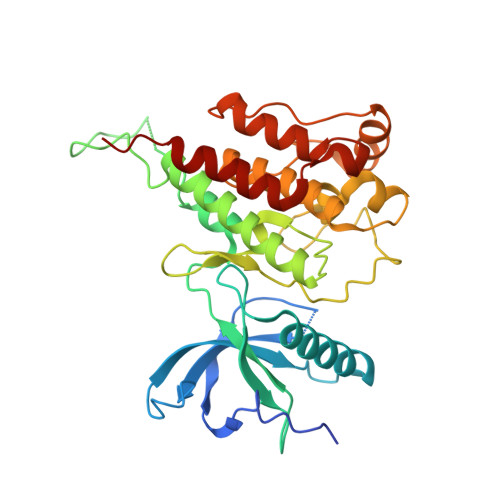 | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for P22455 (Homo sapiens) Explore P22455 Go to UniProtKB: P22455 | |||||
PHAROS: P22455 GTEx: ENSG00000160867 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P22455 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| GX3 (Subject of Investigation/LOI) Query on GX3 | B [auth A] | 6-[2,6-bis(chloranyl)-3,5-dimethoxy-phenyl]-2-(methylamino)-8-[3-(4-prop-2-enoylpiperazin-1-yl)propyl]pyrido[2,3-d]pyrimidin-7-one C26 H30 Cl2 N6 O4 PUIXMSRTTHLNKI-UHFFFAOYSA-N |  | ||
| SO4 Query on SO4 | C [auth A], D [auth A], E [auth A] | SULFATE ION O4 S QAOWNCQODCNURD-UHFFFAOYSA-L |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 42.449 | α = 90 |
| b = 61.1 | β = 97.65 |
| c = 61.041 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| HKL-3000 | data scaling |
| PDB_EXTRACT | data extraction |
| HKL-3000 | data reduction |
| PHENIX | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Natural Science Foundation of China (NSFC) | China | 81570537 |
| National Natural Science Foundation of China (NSFC) | China | 81974074 |
| National Natural Science Foundation of China (NSFC) | China | 31900880 |