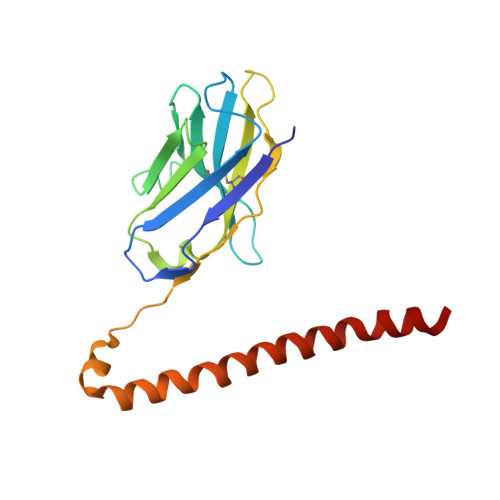
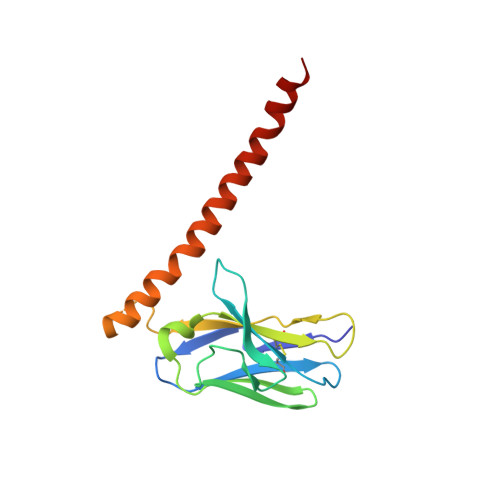
Structural mechanism of laminin recognition by integrin.
Arimori, T., Miyazaki, N., Mihara, E., Takizawa, M., Taniguchi, Y., Cabanas, C., Sekiguchi, K., Takagi, J.(2021) Nat Commun 12: 4012-4012
- PubMed: 34188035
- DOI: https://doi.org/10.1038/s41467-021-24184-8
- Primary Citation of Related Structures:
7CEA, 7CEB, 7CEC - PubMed Abstract:
Recognition of laminin by integrin receptors is central to the epithelial cell adhesion to basement membrane, but the structural background of this molecular interaction remained elusive. Here, we report the structures of the prototypic laminin receptor α6β1 integrin alone and in complex with three-chain laminin-511 fragment determined via crystallography and cryo-electron microscopy, respectively. The laminin-integrin interface is made up of several binding sites located on all five subunits, with the laminin γ1 chain C-terminal portion providing focal interaction using two carboxylate anchor points to bridge metal-ion dependent adhesion site of integrin β1 subunit and Asn189 of integrin α6 subunit. Laminin α5 chain also contributes to the affinity and specificity by making electrostatic interactions with large surface on the β-propeller domain of α6, part of which comprises an alternatively spliced X1 region. The propeller sheet corresponding to this region shows unusually high mobility, suggesting its unique role in ligand capture.
Organizational Affiliation:
Laboratory for Protein Synthesis and Expression, Institute for Protein Research, Osaka University, Suita, Osaka, Japan.