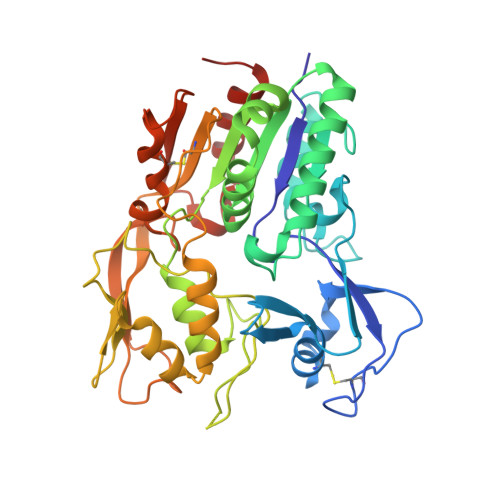
Molecular basis for activation of lecithin:cholesterol acyltransferase by a compound that increases HDL cholesterol.
Manthei, K.A., Yang, S.M., Baljinnyam, B., Chang, L., Glukhova, A., Yuan, W., Freeman, L.A., Maloney, D.J., Schwendeman, A., Remaley, A.T., Jadhav, A., Tesmer, J.J.(2018) Elife 7
- PubMed: 30479275
- DOI: https://doi.org/10.7554/eLife.41604
- Primary Citation of Related Structures:
6MVD - PubMed Abstract:
Lecithin:cholesterol acyltransferase (LCAT) and LCAT-activating compounds are being investigated as treatments for coronary heart disease (CHD) and familial LCAT deficiency (FLD). Herein we report the crystal structure of human LCAT in complex with a potent piperidinylpyrazolopyridine activator and an acyl intermediate-like inhibitor, revealing LCAT in an active conformation. Unlike other LCAT activators, the piperidinylpyrazolopyridine activator binds exclusively to the membrane-binding domain (MBD). Functional studies indicate that the compound does not modulate the affinity of LCAT for HDL, but instead stabilizes residues in the MBD and facilitates channeling of substrates into the active site. By demonstrating that these activators increase the activity of an FLD variant, we show that compounds targeting the MBD have therapeutic potential. Our data better define the substrate binding site of LCAT and pave the way for rational design of LCAT agonists and improved biotherapeutics for augmenting or restoring reverse cholesterol transport in CHD and FLD patients.
Organizational Affiliation:
Life Sciences Institute, University of Michigan, Ann Arbor, United States.