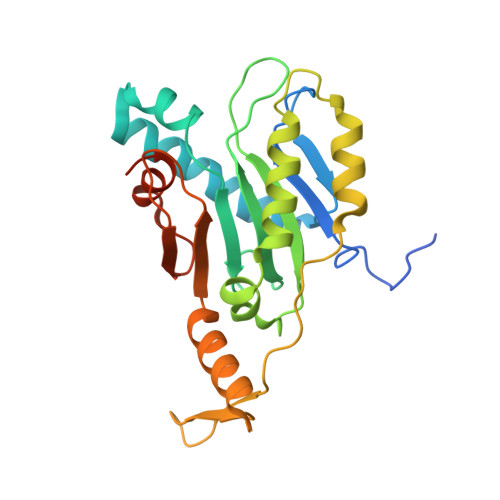
Crystal Structure of the Type VI Secretion System Accessory Protein TagF from Pseudomonas Aeruginosa.
Ok, C.K., Chang, J.H.(2019) Protein Pept Lett 26: 204-214
- PubMed: 30659530
- DOI: https://doi.org/10.2174/0929866526666190119121859
- Primary Citation of Related Structures:
6IZ8 - PubMed Abstract:
Type VI Secretion System (T6SS) has been found in approximately onequarter of the gram-negative bacterial species, and its structural characteristics appear to slightly differ from species to species. The genes encoding T6SS are designated as type six secretion A-M (tssA-M). The expression of the tss gene cluster is regulated by various accessory genes, designated as type VI-associated genes A-P (tagA-P). Tag family proteins have been commonly found in bacteria expressing T6SS but not in all bacterial species. For instance, the tag gene cluster is well-conserved in Pseudomonas aeruginosa (Pa). The PaTagF protein has large homology with ImpM in Rhizobium leguminosarum and SciT in Salmonella enterica. The overexpression of PaTagF represses T6SS complex accumulation and suppresses T6SS antibacterial activity. Thus, the functions of TagF are mediated through direct interactions with the forkhead-associated protein Fha, as evident from the results of the yeast-two hybrid assays. Fha is involved in recruiting a membrane-associated complex either in threonine phosphorylation pathway-dependent or - independent manner. However, functional reports of the tag gene cluster are still limited. In this article, our motivation is to understand the molecular mechanism underlying the regulation of expression of the type VI secretion system complex. In this article, we start with obtaining the gene encoding PaTagF protein by polymerase chain reaction (PCR). Subsequently, the cloned gene is applied to overexpress of PaTagF protein in Escherichia coli, then purify the recombinant PaTagF protein. Thereafter, the protein is crystallized in a condition of 2.5 M NaCl, 0.1 M imidazole (pH 8.0), 3.2 M NaCl, 0.1 M BIS-TRIS propane (pH 7.0) and diffraction datasets of the PaTagF crystals are collected at the Pohang Accelerator Laboratory (PAL). The molecular structure of PaTagF protein is determined by molecular replacement using the uncharacterized protein PA0076 (PDB code:2QNU) as an initial search model by PHENIX crystallographic software package. Model building of PaTagF structure is performed using Coot program. Finally, the structural model is validated using phenix.refine program. PaTagF exists as a tetramer in the asymmetric unit, and the overall fold of each monomer is composed of continuous beta-sheets wrapped by alpha-helices. Each monomer has variable conformations and lengths of both the N- and C-termini. Twelve residues, including the His6 tag from the N-terminus of a symmetry-related molecule, have been found in two of the tetrameric PaTagF structures. A structural homology search revealed that PaTagF was similar to the α-β-α sandwichlike structure of the longin domain on the differentially expressed in normal and neoplastic (DENN) superfamily, which is commonly found in proteins related to trafficking. The tetrameric structure of PaTagF comprises varied N- and C-terminal regions in each subunit and may be stabilized by a symmetry-related molecule. This feature was also shown in the TssL structure from V. cholerae. Furthermore, our study showed that the overall fold of PaTagF is homologous to the longin domain of the DENN family. Therefore, further studies are warranted to elucidate the structure-based evolutionary relationship between protein transport systems from the bacteria and eukaryotic cells.
Organizational Affiliation:
Department of Biology Education, Kyungpook National University, Daegu 41566, South Korea.