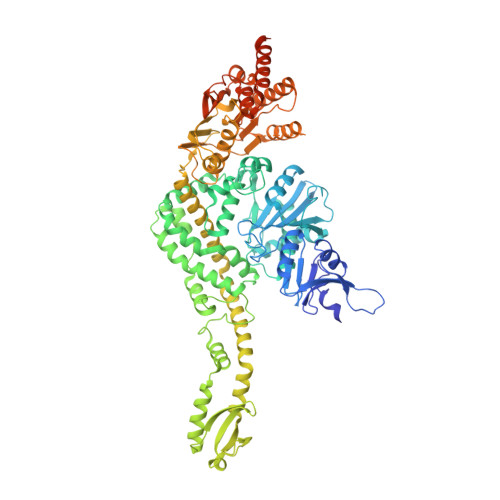
Sharp kinking of a coiled-coil in MutS allows DNA binding and release.
Bhairosing-Kok, D., Groothuizen, F.S., Fish, A., Dharadhar, S., Winterwerp, H.H.K., Sixma, T.K.(2019) Nucleic Acids Res 47: 8888-8898
- PubMed: 31372631
- DOI: https://doi.org/10.1093/nar/gkz649
- Primary Citation of Related Structures:
6I5F - PubMed Abstract:
DNA mismatch repair (MMR) corrects mismatches, small insertions and deletions in DNA during DNA replication. While scanning for mismatches, dimers of MutS embrace the DNA helix with their lever and clamp domains. Previous studies indicated generic flexibility of the lever and clamp domains of MutS prior to DNA binding, but whether this was important for MutS function was unknown. Here, we present a novel crystal structure of DNA-free Escherichia coli MutS. In this apo-structure, the clamp domains are repositioned due to kinking at specific sites in the coiled-coil region in the lever domains, suggesting a defined hinge point. We made mutations at the coiled-coil hinge point. The mutants made to disrupt the helical fold at the kink site diminish DNA binding, whereas those made to increase stability of coiled-coil result in stronger DNA binding. These data suggest that the site-specific kinking of the coiled-coil in the lever domain is important for loading of this ABC-ATPase on DNA.
Organizational Affiliation:
Division of Biochemistry and Oncode Institute, Netherlands Cancer Institute, 1066 CX Amsterdam, the Netherlands.