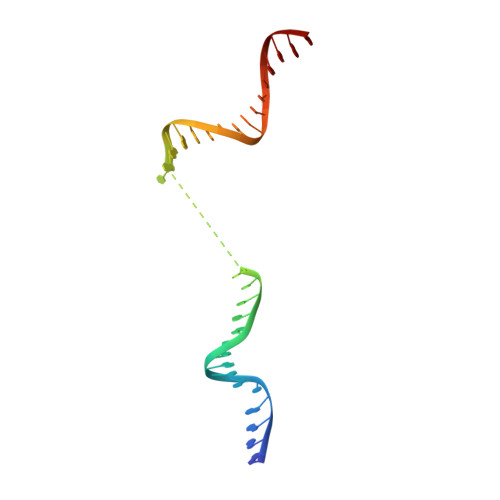
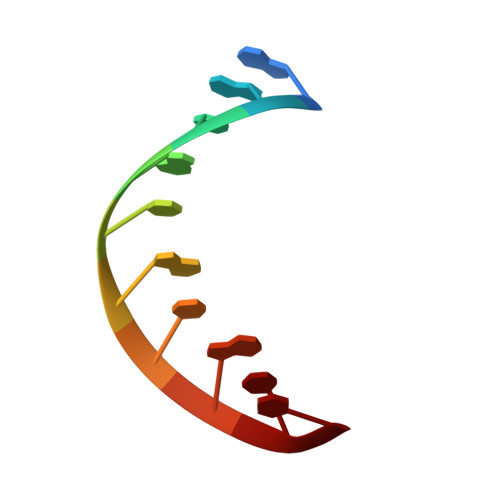
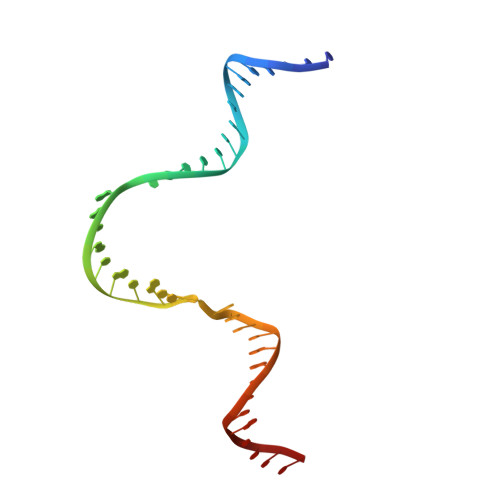
Structural Basis for NusA Stabilized Transcriptional Pausing.
Guo, X., Myasnikov, A.G., Chen, J., Crucifix, C., Papai, G., Takacs, M., Schultz, P., Weixlbaumer, A.(2018) Mol Cell 69: 816-827.e4
- PubMed: 29499136
- DOI: https://doi.org/10.1016/j.molcel.2018.02.008
- Primary Citation of Related Structures:
6FLP, 6FLQ - PubMed Abstract:
Transcriptional pausing by RNA polymerases (RNAPs) is a key mechanism to regulate gene expression in all kingdoms of life and is a prerequisite for transcription termination. The essential bacterial transcription factor NusA stimulates both pausing and termination of transcription, thus playing a central role. Here, we report single-particle electron cryo-microscopy reconstructions of NusA bound to paused E. coli RNAP elongation complexes with and without a pause-enhancing hairpin in the RNA exit channel. The structures reveal four interactions between NusA and RNAP that suggest how NusA stimulates RNA folding, pausing, and termination. An asymmetric translocation intermediate of RNA and DNA converts the active site of the enzyme into an inactive state, providing a structural explanation for the inhibition of catalysis. Comparing RNAP at different stages of pausing provides insights on the dynamic nature of the process and the role of NusA as a regulatory factor.
Organizational Affiliation:
Department of Integrated Structural Biology, Institut de Génétique et de Biologie Moléculaire et Cellulaire (IGBMC), 67404 Illkirch Cedex, France; Université de Strasbourg, 67404 Illkirch Cedex, France; Centre National de la Recherche Scientifique (CNRS), UMR 7104, 67404 Illkirch Cedex, France; Institut National de la Santé et de la Recherche Médicale (Inserm), U964, 67404 Illkirch Cedex, France.