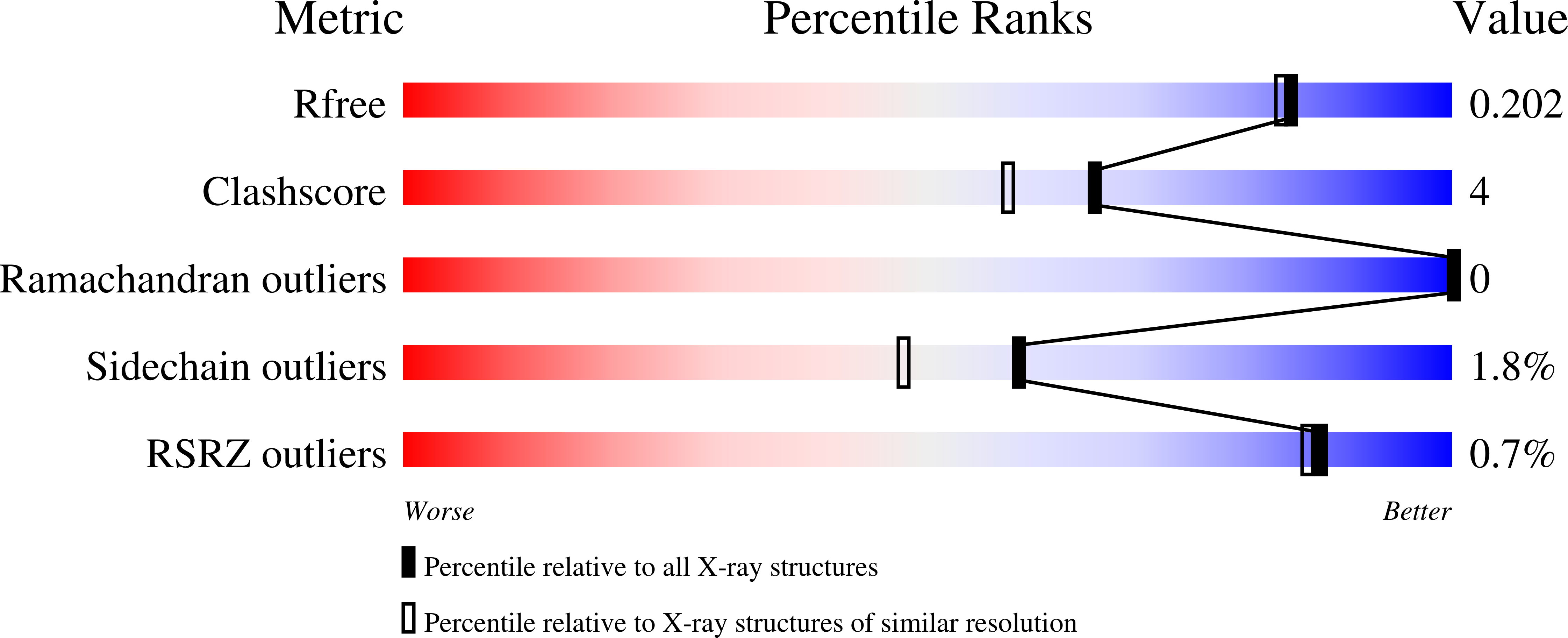
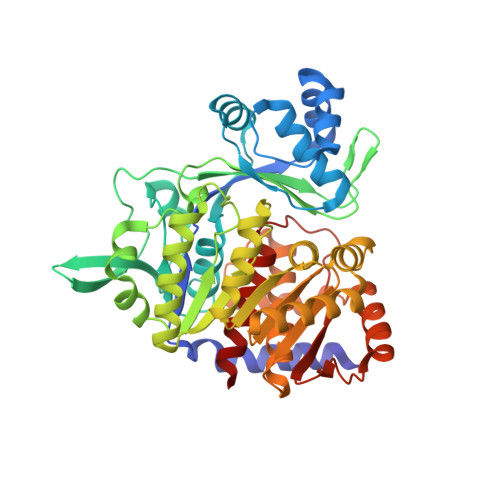
Structural basis for ADP-dependent glucokinase inhibition by 8-bromo-substituted adenosine nucleotide.
Grudnik, P., Kaminski, M.M., Rembacz, K.P., Kuska, K., Madej, M., Potempa, J., Dawidowski, M., Dubin, G.(2018) J Biol Chem 293: 11088-11099
- PubMed: 29784881
- DOI: https://doi.org/10.1074/jbc.RA117.001562
- Primary Citation of Related Structures:
5O0I, 5O0J - PubMed Abstract:
In higher eukaryotes, several ATP-utilizing enzymes known as hexokinases activate glucose in the glycolysis pathway by phosphorylation to glucose 6-phosphate. In contrast to canonical hexokinases, which use ATP, ADP-dependent glucokinase (ADPGK) catalyzes noncanonical phosphorylation of glucose to glucose 6-phosphate using ADP as a phosphate donor. Initially discovered in Archaea, the human homolog of ADPGK was described only recently. ADPGK's involvement in modified bioenergetics of activated T cells has been postulated, and elevated ADPGK expression has been reported in various cancer tissues. However, the physiological role of ADPGK is still poorly understood, and effective ADPGK inhibitors still await discovery. Here, we show that 8-bromo-substituted adenosine nucleotide inhibits human ADPGK. By solving the crystal structure of archaeal ADPGK in complex with 8-bromoadenosine phosphate (8-Br-AMP) at 1.81 Å resolution, we identified the mechanism of inhibition. We observed that 8-Br-AMP is a competitive inhibitor of ADPGK and that the bromine substitution induces marked structural changes within the protein's active site by engaging crucial catalytic residues. The results obtained using the Jurkat model of activated human T cells suggest its moderate activity in a cellular setting. We propose that our structural insights provide a critical basis for rational development of novel ADPGK inhibitors.
Organizational Affiliation:
From the Faculty of Biochemistry, Biophysics and Biotechnology and przemyslaw.grudnik@uj.edu.pl.