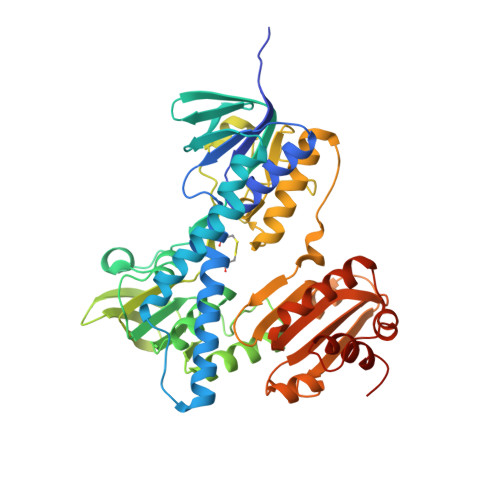
Biochemical and Structural Properties of a Thermostable Mercuric Ion Reductase from Metallosphaera sedula.
Artz, J.H., White, S.N., Zadvornyy, O.A., Fugate, C.J., Hicks, D., Gauss, G.H., Posewitz, M.C., Boyd, E.S., Peters, J.W.(2015) Front Bioeng Biotechnol 3: 97-97
- PubMed: 26217660
- DOI: https://doi.org/10.3389/fbioe.2015.00097
- Primary Citation of Related Structures:
4YWO - PubMed Abstract:
Mercuric ion reductase (MerA), a mercury detoxification enzyme, has been tuned by evolution to have high specificity for mercuric ions (Hg(2+)) and to catalyze their reduction to a more volatile, less toxic elemental form. Here, we present a biochemical and structural characterization of MerA from the thermophilic crenarchaeon Metallosphaera sedula. MerA from M. sedula is a thermostable enzyme, and remains active after extended incubation at 97°C. At 37°C, the NADPH oxidation-linked Hg(2+) reduction specific activity was found to be 1.9 μmol/min⋅mg, increasing to 3.1 μmol/min⋅mg at 70°C. M. sedula MerA crystals were obtained and the structure was solved to 1.6 Å, representing the first solved crystal structure of a thermophilic MerA. Comparison of both the crystal structure and amino acid sequence of MerA from M. sedula to mesophillic counterparts provides new insights into the structural determinants that underpin the thermal stability of the enzyme.
Organizational Affiliation:
Department of Chemistry and Biochemistry, Montana State University , Bozeman, MT , USA.