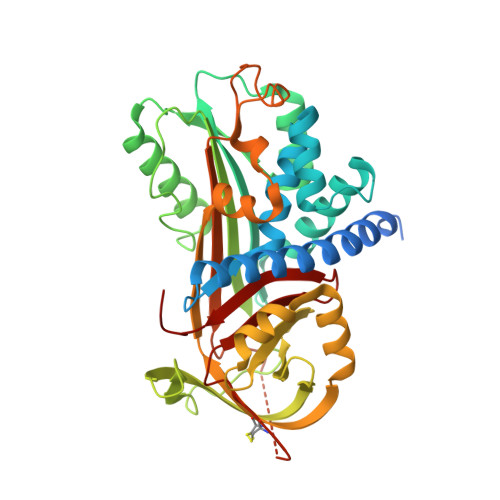
A unique serpin P1' glutamate and a conserved beta-sheet C arginine are key residues for activity, protease recognition and stability of serpinA12 (vaspin).
Ulbricht, D., Pippel, J., Schultz, S., Meier, R., Strater, N., Heiker, J.T.(2015) Biochem J 470: 357-367
- PubMed: 26199422
- DOI: https://doi.org/10.1042/BJ20150643
- Primary Citation of Related Structures:
4Y3K, 4Y40 - PubMed Abstract:
SerpinA12 (vaspin) is thought to be mainly expressed in adipose tissue and has multiple beneficial effects on metabolic, inflammatory and atherogenic processes related to obesity. KLK7 (kallikrein 7) is the only known protease target of vaspin to date and is inhibited with a moderate inhibition rate. In the crystal structure, the cleavage site (P1-P1') of the vaspin reactive centre loop is fairly rigid compared with the flexible residues before P2, possibly supported by an ionic interaction of P1' glutamate (Glu(379)) with an arginine residue (Arg(302)) of the β-sheet C. A P1' glutamate seems highly unusual and unfavourable for the protease KLK7. We characterized vaspin mutants to investigate the roles of these two residues in protease inhibition and recognition by vaspin. Reactive centre loop mutations changing the P1' residue or altering the reactive centre loop conformation significantly increased inhibition parameters, whereas removal of the positive charge within β-sheet C impeded the serpin-protease interaction. Arg(302) is a crucial contact to enable vaspin recognition by KLK7 and it supports moderate inhibition of the serpin despite the presence of the detrimental P1' Glu(379), which clearly represents a major limiting factor for vaspin-inhibitory activity. We also show that the vaspin-inhibition rate for KLK7 can be modestly increased by heparin and demonstrate that vaspin is a heparin-binding serpin. Noteworthily, we observed vaspin as a remarkably thermostable serpin and found that Glu(379) and Arg(302) influence heat-induced polymerization. These structural and functional results reveal the mechanistic basis of how reactive centre loop sequence and exosite interaction in vaspin enable KLK7 recognition and regulate protease inhibition as well as stability of this adipose tissue-derived serpin.
Organizational Affiliation:
Institute of Biochemistry, Faculty of Biosciences, Pharmacy and Psychology, University of Leipzig, 04103 Leipzig, Germany.