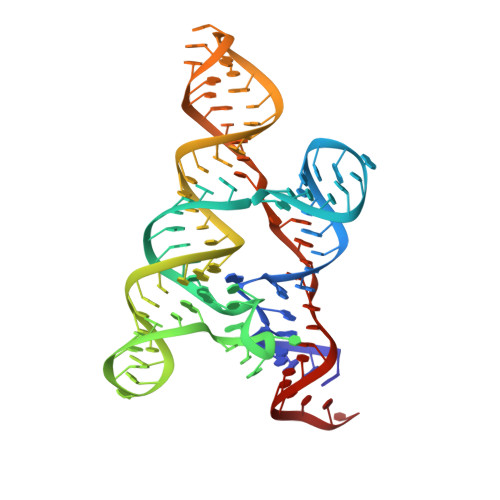
Mn(2+)-Sensing Mechanisms of yybP-ykoY Orphan Riboswitches.
Price, I.R., Gaballa, A., Ding, F., Helmann, J.D., Ke, A.(2015) Mol Cell 57: 1110-1123
- PubMed: 25794619
- DOI: https://doi.org/10.1016/j.molcel.2015.02.016
- Primary Citation of Related Structures:
4Y1I, 4Y1J, 4Y1M - PubMed Abstract:
Gene regulation in cis by riboswitches is prevalent in bacteria. The yybP-ykoY riboswitch family is quite widespread, yet its ligand and function remained unknown. Here, we characterize the Lactococcus lactis yybP-ykoY orphan riboswitch as a Mn(2+)-dependent transcription-ON riboswitch, with a ∼30-40 μM affinity for Mn(2+). We further determined its crystal structure at 2.7 Å to elucidate the metal sensing mechanism. The riboswitch resembles a hairpin, with two coaxially stacked helices tethered by a four-way junction and a tertiary docking interface. The Mn(2+)-sensing region, strategically located at the highly conserved docking interface, has two metal binding sites. Whereas one site tolerates the binding of either Mg(2+) or Mn(2+), the other site strongly prefers Mn(2+) due to a direct contact from the N7 of an invariable adenosine. Mutagenesis and a Mn(2+)-free E. coli yybP-ykoY structure further reveal that Mn(2+) binding is coupled with stabilization of the Mn(2+)-sensing region and the aptamer domain.
Organizational Affiliation:
Department of Molecular Biology and Genetics, Cornell University, 253 Biotechnology Building, Ithaca, NY 14853, USA.