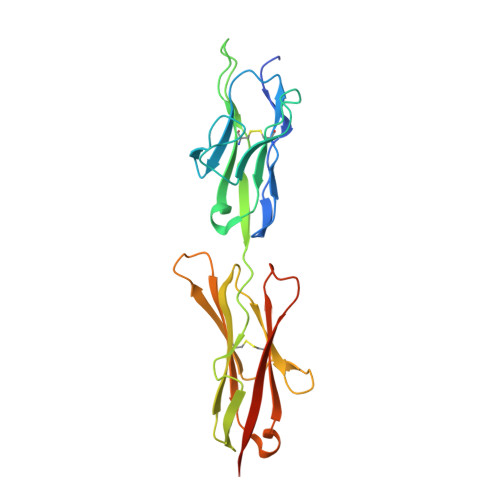
Extracellular Architecture of the SYG-1/SYG-2 Adhesion Complex Instructs Synaptogenesis.
Ozkan, E., Chia, P.H., Wang, R.R., Goriatcheva, N., Borek, D., Otwinowski, Z., Walz, T., Shen, K., Garcia, K.C.(2014) Cell 156: 482-494
- PubMed: 24485456
- DOI: https://doi.org/10.1016/j.cell.2014.01.004
- Primary Citation of Related Structures:
4OF0, 4OF3, 4OF6, 4OF7, 4OF8, 4OFD, 4OFI, 4OFK, 4OFP, 4OFY - PubMed Abstract:
SYG-1 and SYG-2 are multipurpose cell adhesion molecules (CAMs) that have evolved across all major animal taxa to participate in diverse physiological functions, ranging from synapse formation to formation of the kidney filtration barrier. In the crystal structures of several SYG-1 and SYG-2 orthologs and their complexes, we find that SYG-1 orthologs homodimerize through a common, bispecific interface that similarly mediates an unusual orthogonal docking geometry in the heterophilic SYG-1/SYG-2 complex. C. elegans SYG-1's specification of proper synapse formation in vivo closely correlates with the heterophilic complex affinity, which appears to be tuned for optimal function. Furthermore, replacement of the interacting domains of SYG-1 and SYG-2 with those from CAM complexes that assume alternative docking geometries or the introduction of segmental flexibility compromised synaptic function. These results suggest that SYG extracellular complexes do not simply act as "molecular velcro" and that their distinct structural features are important in instructing synaptogenesis. PAPERFLICK:
- Department of Molecular and Cellular Physiology and Department of Structural Biology, Stanford University School of Medicine, Stanford, CA 94305, USA; Howard Hughes Medical Institute, Stanford University School of Medicine, Stanford, CA 94305, USA.
Organizational Affiliation: