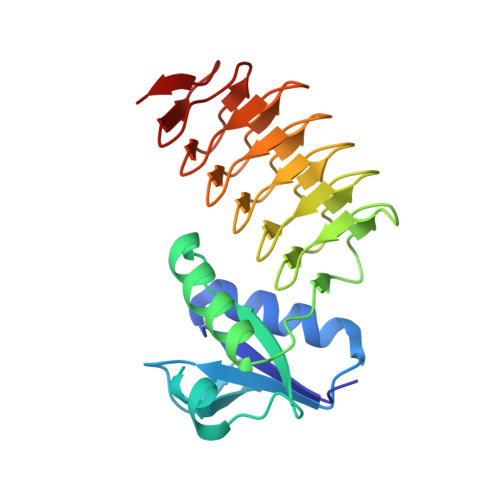
Biochemical analysis and structure determination of bacterial acetyltransferases responsible for the biosynthesis of UDP-N,N'-diacetylbacillosamine.
Morrison, M.J., Imperiali, B.(2013) J Biol Chem 288: 32248-32260
- PubMed: 24064219
- DOI: https://doi.org/10.1074/jbc.M113.510560
- Primary Citation of Related Structures:
4M98, 4M99, 4M9C - PubMed Abstract:
UDP-N,N'-diacetylbacillosamine (UDP-diNAcBac) is a unique carbohydrate produced by a number of bacterial species and has been implicated in pathogenesis. The terminal step in the formation of this important bacterial sugar is catalyzed by an acetyl-CoA (AcCoA)-dependent acetyltransferase in both N- and O-linked protein glycosylation pathways. This bacterial acetyltransferase is a member of the left-handed β-helix family and forms a homotrimer as the functional unit. Whereas previous endeavors have focused on the Campylobacter jejuni acetyltransferase (PglD) from the N-linked glycosylation pathway, structural characterization of the homologous enzymes in the O-linked glycosylation pathways is lacking. Herein, we present the apo-crystal structures of the acetyltransferase domain (ATD) from the bifunctional enzyme PglB (Neisseria gonorrhoeae) and the full-length acetyltransferase WeeI (Acinetobacter baumannii). Additionally, a PglB-ATD structure was solved in complex with AcCoA. Surprisingly, this structure reveals a contrasting binding mechanism for this substrate when compared with the AcCoA-bound PglD structure. A comparison between these findings and the previously solved PglD crystal structures illustrates a dichotomy among N- and O-linked glycosylation pathway enzymes. Based upon these structures, key residues in the UDP-4-amino and AcCoA binding pockets were mutated to determine their effect on binding and catalysis in PglD, PglB-ATD, and WeeI. Last, a phylogenetic analysis of the aforementioned acetyltransferases was employed to illuminate the diversity among N- and O-linked glycosylation pathway enzymes.
Organizational Affiliation:
From the Departments of Chemistry and Biology, Massachusetts Institute of Technology, Cambridge, Massachusetts 02139.