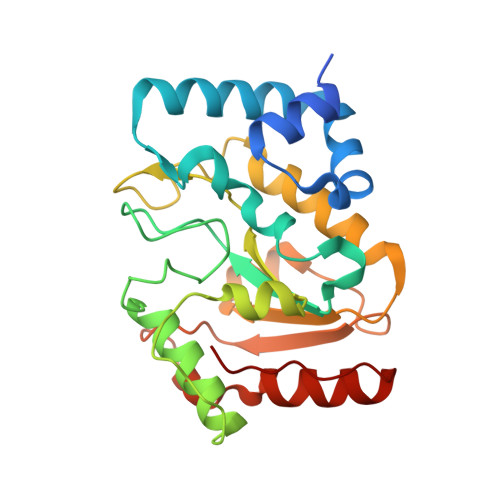
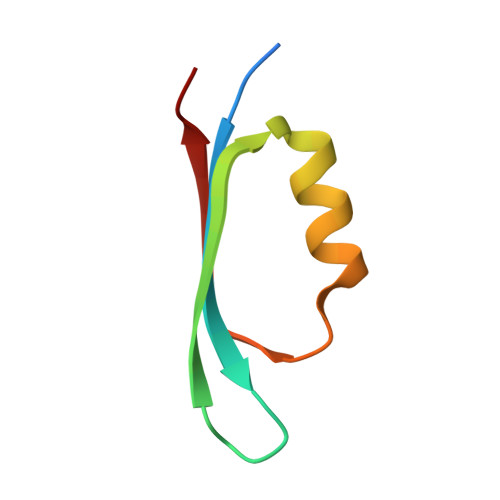
Architecturally diverse proteins converge on an analogous mechanism to inactivate Uracil-DNA glycosylase.
Cole, A.R., Ofer, S., Ryzhenkova, K., Baltulionis, G., Hornyak, P., Savva, R.(2013) Nucleic Acids Res 41: 8760-8775
- PubMed: 23892286
- DOI: https://doi.org/10.1093/nar/gkt633
- Primary Citation of Related Structures:
4L5N - PubMed Abstract:
Uracil-DNA glycosylase (UDG) compromises the replication strategies of diverse viruses from unrelated lineages. Virally encoded proteins therefore exist to limit, inhibit or target UDG activity for proteolysis. Viral proteins targeting UDG, such as the bacteriophage proteins ugi, and p56, and the HIV-1 protein Vpr, share no sequence similarity, and are not structurally homologous. Such diversity has hindered identification of known or expected UDG-inhibitory activities in other genomes. The structural basis for UDG inhibition by ugi is well characterized; yet, paradoxically, the structure of the unbound p56 protein is enigmatically unrevealing of its mechanism. To resolve this conundrum, we determined the structure of a p56 dimer bound to UDG. A helix from one of the subunits of p56 occupies the UDG DNA-binding cleft, whereas the dimer interface forms a hydrophobic box to trap a mechanistically important UDG residue. Surprisingly, these p56 inhibitory elements are unexpectedly analogous to features used by ugi despite profound architectural disparity. Contacts from B-DNA to UDG are mimicked by residues of the p56 helix, echoing the role of ugi's inhibitory beta strand. Using mutagenesis, we propose that DNA mimicry by p56 is a targeting and specificity mechanism supporting tight inhibition via hydrophobic sequestration.
Organizational Affiliation:
Department of Biological Sciences, Institute of Structural and Molecular Biology, Birkbeck College, University of London, Malet Street, London WC1E 7HX, UK and Research Department of Structural and Molecular Biology, Institute of Structural and Molecular Biology, University College London, Gower Street, London WC1E 6BT, UK.