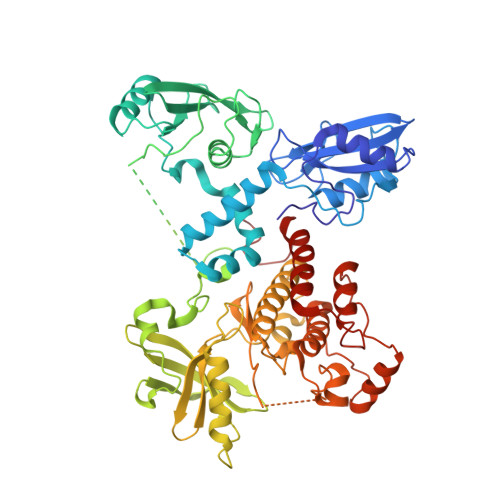
Structural Basis for Activation of ZAP-70 by Phosphorylation of the SH2-Kinase Linker.
Yan, Q., Barros, T., Visperas, P.R., Deindl, S., Kadlecek, T.A., Weiss, A., Kuriyan, J.(2013) Mol Cell Biol 33: 2188-2201
- PubMed: 23530057
- DOI: https://doi.org/10.1128/MCB.01637-12
- Primary Citation of Related Structures:
4K2R - PubMed Abstract:
Serial activation of the tyrosine kinases Lck and ZAP-70 initiates signaling downstream of the T cell receptor. We previously reported the structure of an autoinhibited ZAP-70 variant in which two regulatory tyrosine residues (315 and 319) in the SH2-kinase linker were replaced by phenylalanine. We now present a crystal structure of ZAP-70 in which Tyr 315 and Tyr 319 are not mutated, leading to the recognition of a five-residue sequence register error in the SH2-kinase linker of the original crystallographic model. The revised model identifies distinct roles for these two tyrosines. As seen in a recently reported structure of the related tyrosine kinase Syk, Tyr 315 of ZAP-70 is part of a hydrophobic interface between the regulatory apparatus and the kinase domain, and the integrity of this interface would be lost upon engagement of doubly phosphorylated peptides by the SH2 domains. Tyr 319 is not necessarily dislodged by SH2 engagement, which activates ZAP-70 only ∼5-fold in vitro. In contrast, phosphorylation by Lck activates ZAP-70 ∼100-fold. This difference is due to the ability of Tyr 319 to suppress ZAP-70 activity even when the SH2 domains are dislodged from the kinase domain, providing stringent control of ZAP-70 activity downstream of Lck.
Organizational Affiliation:
Department of Molecular and Cell Biology, University of California, Berkeley, CA, USA.