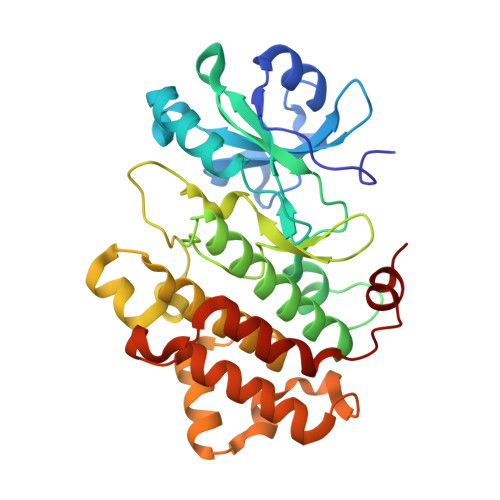
Structural Characterization of the RLCK Family Member BSK8: A Pseudokinase with an Unprecedented Architecture
Grutter, C., Sreeramulu, S., Sessa, G., Rauh, D.(2013) J Mol Biol 425: 4455-4467
- PubMed: 23911552
- DOI: https://doi.org/10.1016/j.jmb.2013.07.034
- Primary Citation of Related Structures:
4I92, 4I93, 4I94 - PubMed Abstract:
Brassinosteroid signaling kinases (BSKs) are plant-specific receptor-like cytoplasmic protein kinases involved in the brassinosteroid signaling pathway. Unlike common protein kinases, they possess a naturally occurring alanine residue at the "gatekeeper" position, as well as other sequence variations. How BSKs activate downstream proteins such as BSU1, as well as the structural consequences of their unusual sequential features, was unclear. We crystallized the catalytic domain of BSK8 and solved its structure by multiple-wavelength anomalous dispersion phasing methods to a resolution of 1.5Å. In addition, a co-crystal structure of BSK8 with 5-adenylyl imidodiphosphate (AMP-PNP) revealed unusual conformational arrangements of the nucleotide phosphate groups and catalytic key motifs, typically not observed for active protein kinases. Sequential analysis and comparisons with known pseudokinase structures suggest that BSKs represent constitutively inactive protein kinases that regulate brassinosteroid signal transfer through an allosteric mechanism.
Organizational Affiliation:
Technische Universität Dortmund Fakultät Chemie und Chemische Biologie, Otto-Hahn-Strasse 6, D-44227 Dortmund, Germany.