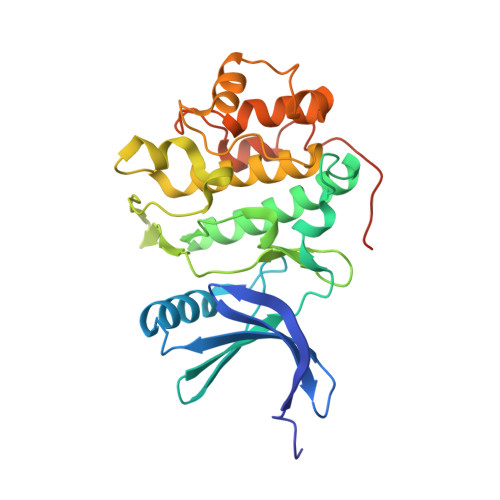
Structure-based design and optimization of 2-aminothiazole-4-carboxamide as a new class of CHK1 inhibitors.
Huang, X., Cheng, C.C., Fischmann, T.O., Duca, J.S., Richards, M., Tadikonda, P.K., Reddy, P.A., Zhao, L., Arshad Siddiqui, M., Parry, D., Davis, N., Seghezzi, W., Wiswell, D., Shipps, G.W.(2013) Bioorg Med Chem Lett 23: 2590-2594
- PubMed: 23535330
- DOI: https://doi.org/10.1016/j.bmcl.2013.02.108
- Primary Citation of Related Structures:
4HYH, 4HYI - PubMed Abstract:
Drug design efforts in the emerging 2-aminothiazole-4-carboxamide class of CHK1 inhibitors have uncovered specific combinations of key substructures within the molecule; resulting in significant improvements in cell-based activity while retaining a greater than one hundred-fold selectivity against CDK2. The X-ray crystal structure of a complex between compound 39 and the CHK1 protein detailing a 'U-shaped' topology and key interactions with the protein surface at the ATP site is also reported.
Organizational Affiliation:
Merck Research Laboratories, 320 Bent Street, Cambridge, MA 02141, USA. xiaohua.huang@merck.com