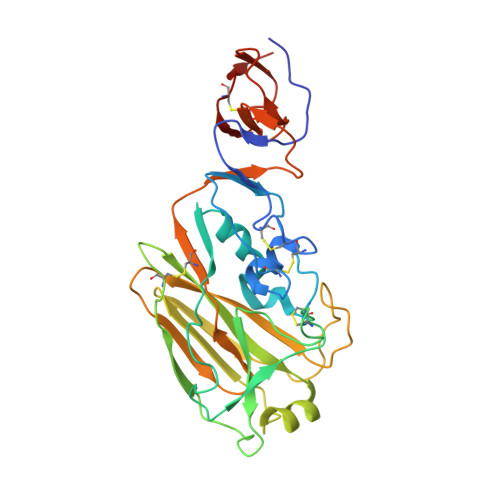
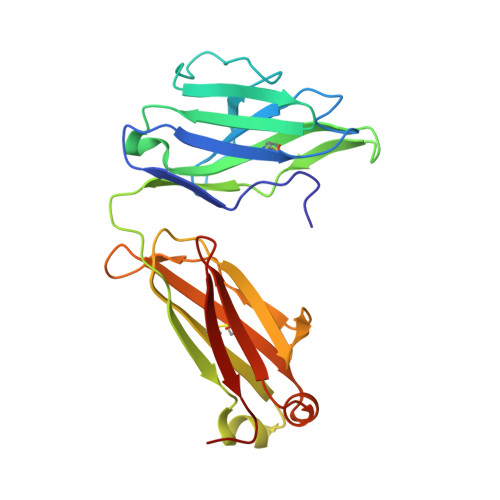
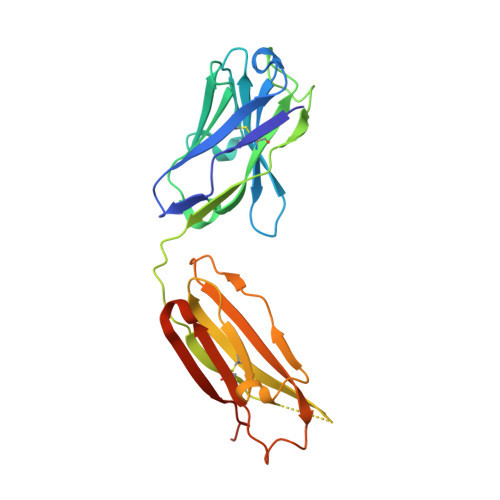
Highly conserved protective epitopes on influenza B viruses.
Dreyfus, C., Laursen, N.S., Kwaks, T., Zuijdgeest, D., Khayat, R., Ekiert, D.C., Lee, J.H., Metlagel, Z., Bujny, M.V., Jongeneelen, M., van der Vlugt, R., Lamrani, M., Korse, H.J., Geelen, E., Sahin, O., Sieuwerts, M., Brakenhoff, J.P., Vogels, R., Li, O.T., Poon, L.L., Peiris, M., Koudstaal, W., Ward, A.B., Wilson, I.A., Goudsmit, J., Friesen, R.H.(2012) Science 337: 1343-1348
- PubMed: 22878502
- DOI: https://doi.org/10.1126/science.1222908
- Primary Citation of Related Structures:
4FQH, 4FQI, 4FQJ, 4FQK, 4FQL, 4FQM, 4FQV, 4FQY - PubMed Abstract:
Identification of broadly neutralizing antibodies against influenza A viruses has raised hopes for the development of monoclonal antibody-based immunotherapy and "universal" vaccines for influenza. However, a substantial part of the annual flu burden is caused by two cocirculating, antigenically distinct lineages of influenza B viruses. Here, we report human monoclonal antibodies, CR8033, CR8071, and CR9114, that protect mice against lethal challenge from both lineages. Antibodies CR8033 and CR8071 recognize distinct conserved epitopes in the head region of the influenza B hemagglutinin (HA), whereas CR9114 binds a conserved epitope in the HA stem and protects against lethal challenge with influenza A and B viruses. These antibodies may inform on development of monoclonal antibody-based treatments and a universal flu vaccine for all influenza A and B viruses.
Organizational Affiliation:
Department of Molecular Biology, The Scripps Research Institute, 10550 North Torrey Pines Road, La Jolla, CA 92037, USA.