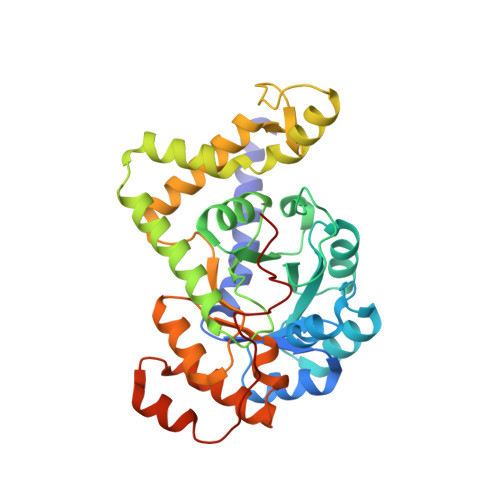
Crystal structure of pyridoxal biosynthesis lyase PdxS from Pyrococcus horikoshii.
Matsuura, A., Yoon, J.Y., Yoon, H.J., Lee, H.H., Suh, S.W.(2012) Mol Cells 34: 407-412
- PubMed: 23104439
- DOI: https://doi.org/10.1007/s10059-012-0198-8
- Primary Citation of Related Structures:
4FIQ, 4FIR - PubMed Abstract:
Pyridoxal 5'-phosphate (PLP) is the biologically active form of vitamin B(6) and is de novo synthesized from three substrates, dihydroxyacetone phosphate (DHAP), riburose 5-phosphate (RBP), and ammonia hydrolysed from glutamine. Glutamine amidotransferase (PdxT) catalyzes the production of ammonia from glutamine, while PdxS catalyzes the following condensation of ribulose 5-phosphate (Ru5P), glyceraldehyde-3-phosphate (G3P), and ammonia. PdxS exists as a hexamer or dodecamer depending on species and makes a 1:1 complex with PdxT. Pyrococcus horikoshii PdxS has a 37 amino acids insertion region, which is found in some archaeal PdxS proteins, but its structure and function are unknown. To provide further structural information on the role of the insertion region, the oligomeric state, and ligand binding mode of P. horikoshii PdxS, the crystal structure of PdxS from P. horikoshii was solved in two forms: (i) apo form, (ii) r ibose 5-phosphate (R5P) complex and the quaternary structure of PdxS in solution was determined by analytical gel filtration. P. horikoshii PdxS forms hexamer in solution based on analytical gel filtration data. When we superimpose the structure of P. horikoshii PdxS with other dodecamer structures of PdxS, the additional insertion is located apart from the active site and induces a steric clash on the hexamer-hexamer interface of PdxS proteins. Our results suggest that the additional insertion perturbs dodecamer formation of P. horikoshii PdxS.
Organizational Affiliation:
Department of Bio & Nano Chemistry, Kookmin University, Seoul 136-702, Korea.