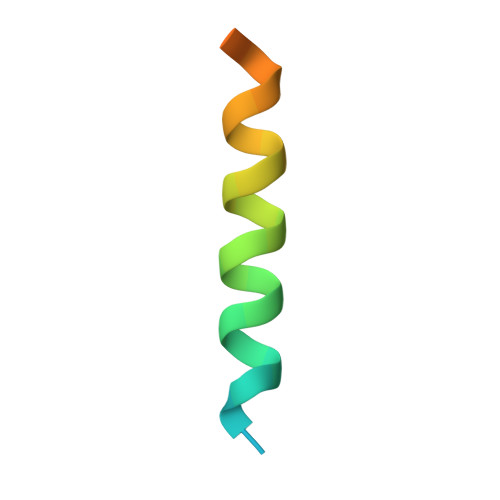
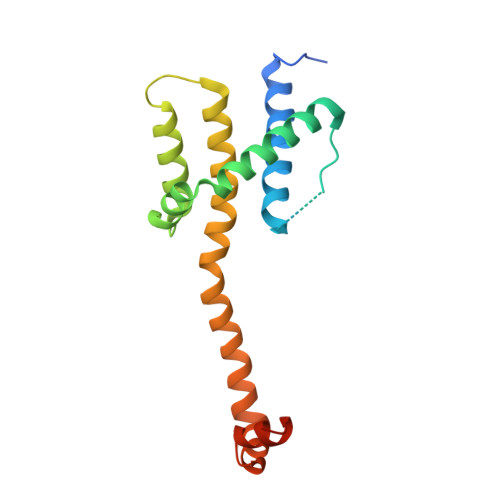
Bax Crystal Structures Reveal How Bh3 Domains Activate Bax and Nucleate its Oligomerization to Induce Apoptosis.
Czabotar, P.E., Westphal, D., Dewson, G., Ma, S., Hockings, C., Fairlie, W.D., Lee, E.F., Yao, S., Robin, A.Y., Smith, B.J., Huang, D.C., Kluck, R.M., Adams, J.M., Colman, P.M.(2013) Cell 152: 519
- PubMed: 23374347
- DOI: https://doi.org/10.1016/j.cell.2012.12.031
- Primary Citation of Related Structures:
4BD2, 4BD6, 4BD7, 4BD8, 4BDU - PubMed Abstract:
In stressed cells, apoptosis ensues when Bcl-2 family members Bax or Bak oligomerize and permeabilize the mitochondrial outer membrane. Certain BH3-only relatives can directly activate them to mediate this pivotal, poorly understood step. To clarify the conformational changes that induce Bax oligomerization, we determined crystal structures of BaxΔC21 treated with detergents and BH3 peptides. The peptides bound the Bax canonical surface groove but, unlike their complexes with prosurvival relatives, dissociated Bax into two domains. The structures define the sequence signature of activator BH3 domains and reveal how they can activate Bax via its groove by favoring release of its BH3 domain. Furthermore, Bax helices α2-α5 alone adopted a symmetric homodimer structure, supporting the proposal that two Bax molecules insert their BH3 domain into each other's surface groove to nucleate oligomerization. A planar lipophilic surface on this homodimer may engage the membrane. Our results thus define critical Bax transitions toward apoptosis.
Organizational Affiliation:
The Walter and Eliza Hall Institute of Medical Research, Melbourne, Victoria 3052, Australia. czabotar@wehi.edu.au