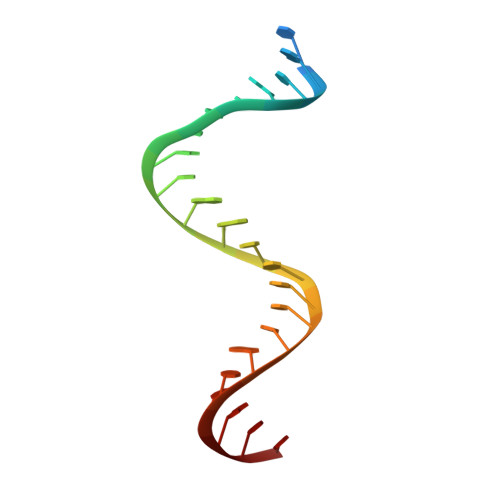
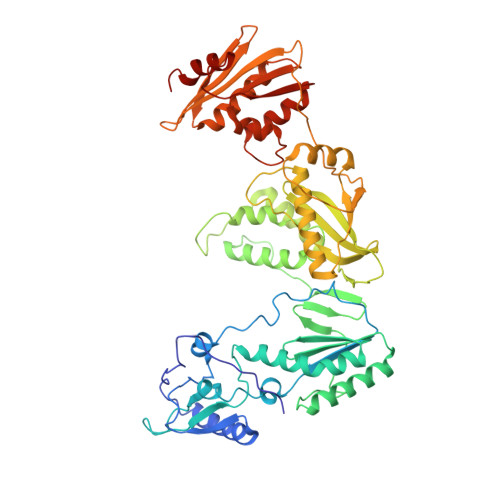
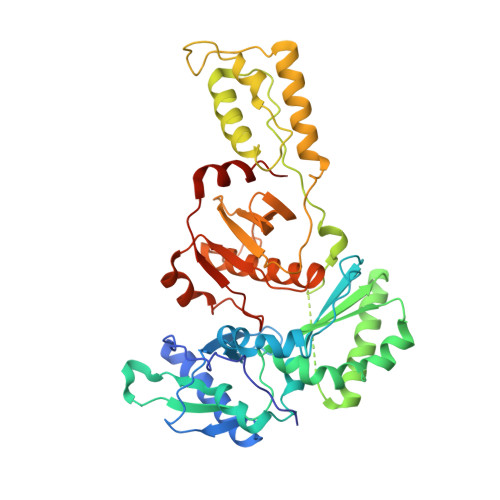
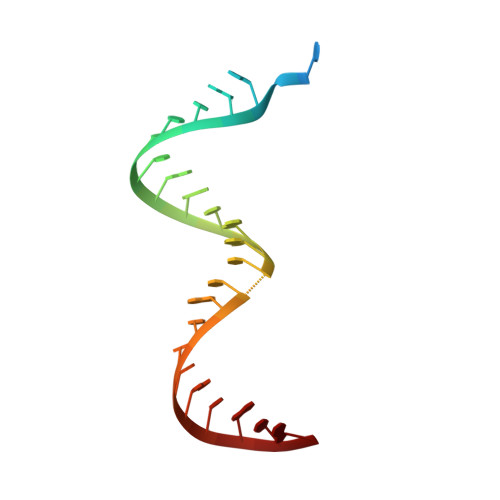
Complexes of HIV-1 RT, Nnrti and RNA/DNA Hybrid Reveal a Structure Compatible with RNA Degradation
Lapkouski, M., Tian, L., Miller, J.T., Le Grice, S.F.J., Yang, W.(2013) Nat Struct Mol Biol 20: 230
- PubMed: 23314251
- DOI: https://doi.org/10.1038/nsmb.2485
- Primary Citation of Related Structures:
4B3O, 4B3P, 4B3Q - PubMed Abstract:
Hundreds of structures of type 1 human immunodeficiency virus (HIV-1) reverse transcriptase (RT) have been determined, but only one contains an RNA/DNA hybrid. Here we report three structures of HIV-1 RT complexed with a non-nucleotide RT inhibitor (NNRTI) and an RNA/DNA hybrid. In the presence of an NNRTI, the RNA/DNA structure differs from all prior nucleic acid-RT structures including the RNA/DNA hybrid. The enzyme structure also differs from all previous RT-DNA complexes. Thus, the hybrid has ready access to the RNase-H active site. These observations indicate that an RT-nucleic acid complex may adopt two structural states, one competent for DNA polymerization and the other for RNA degradation. RT mutations that confer drug resistance but are distant from the inhibitor-binding sites often map to the unique RT-hybrid interface that undergoes conformational changes between two catalytic states.
Organizational Affiliation:
Laboratory of Molecular Biology, National Institute of Diabetes and Digestive and Kidney Diseases, National Institutes of Health, Bethesda, MD 20892, USA.