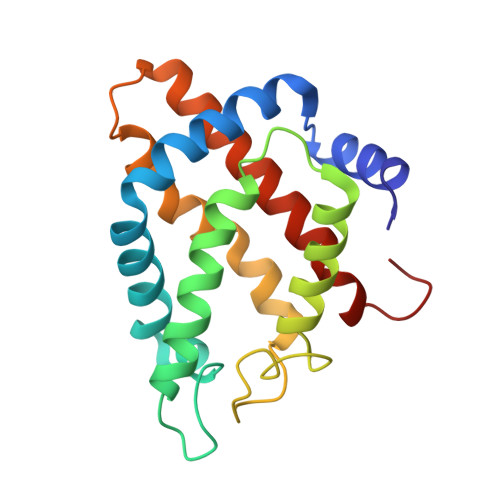
Functional and Structural Role of the N-Terminal Extension in Methanosarcina Acetivorans Protoglobin.
Ciaccio, C., Pesce, A., Tundo, G.R., Tilleman, L., Bertolacci, L., Dewilde, S., Moens, L., Ascenzi, P., Bolognesi, M., Nardini, M., Coletta, M.(2013) Biochim Biophys Acta 1834: 1813
- PubMed: 23485914
- DOI: https://doi.org/10.1016/j.bbapap.2013.02.026
- Primary Citation of Related Structures:
3ZH0 - PubMed Abstract:
Functional and structural properties of protoglobin from Methanosarcina acetivorans, whose Cys(101)E20 residue was mutated to Ser (MaPgb*), and of mutants missing either the first 20 N-terminal amino acids (MaPgb*-ΔN20 mutant), or the first 33 N-terminal amino acids [N-terminal loop of 20 amino acids and a 13-residue Z-helix, preceding the globin fold A-helix; (MaPgb*-ΔN20Z mutant)] have been investigated. In keeping with the MaPgb*-ΔN20 mutant crystal structure, here reported at 2.0Å resolution, which shows an increased exposure of the haem propionates to the solvent, the analysis of ligand binding kinetics highlights high accessibility of ligands to the haem pocket in ferric MaPgb*-ΔN20. CO binding to ferrous MaPgb*-ΔN20 displays a marked biphasic behavior, with a fast binding process close to that observed in MaPgb* and a slow carbonylation process, characterized by a rate-limiting step. Conversely, removal of the first 33 residues induces a substantial perturbation of the overall MaPgb* structure, with loss of α-helical content and potential partial collapse of the protein chain. As such, ligand binding kinetics are characterized by very slow rates that are independent of ligand concentration, this being indicative of a high energy barrier for ligand access to the haem, possibly due to localized misfolding. This article is part of a Special Issue entitled: Oxygen Binding and Sensing Proteins.
Organizational Affiliation:
Department of Clinical Sciences and Translational Medicine, University of Roma Tor Vergata, Roma, Italy.