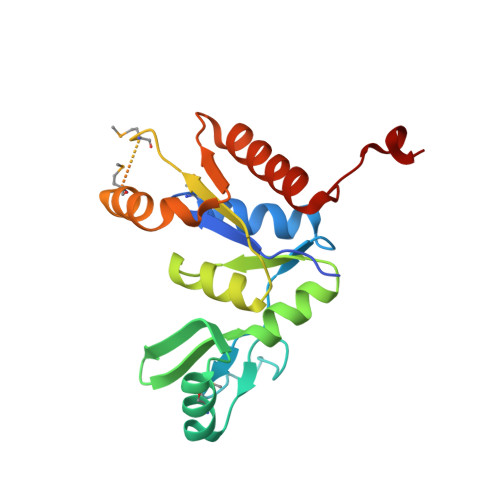
Crystal structure of the guanylate kinase domain from discs large homolog 1 (DLG1/SAP97)
Mori, S., Tezuka, Y., Arakawa, A., Handa, N., Shirouzu, M., Akiyama, T., Yokoyama, S.(2013) Biochem Biophys Res Commun 435: 334-338
- PubMed: 23624197
- DOI: https://doi.org/10.1016/j.bbrc.2013.04.056
- Primary Citation of Related Structures:
3W9Y - PubMed Abstract:
Discs large homolog 1 (DLG1/SAP97) is involved in the development and regulation of neuronal and immunological synapses. DLG1 is a member of the membrane associated guanylate kinase (MAGUK) family of proteins, which function as molecular scaffolds. The C-terminal guanylate kinase (GK) domain of DLG1 binds peptides with a phosphorylated serine residue. In this study, we solved the crystal structure of the GK domain of human DLG1. The C-terminal tail of DLG1 is bound to the peptide-binding site of an adjacent symmetry-related DLG1 GK molecule. The binding direction of the C-terminal tail to the peptide-binding site is opposite to that of the phosphorylated LGN peptide in complex with the rat DLG1 GK domain. The C-terminal tail forms a 310 helix, which is also different from the conformation of the phosphorylated LGN peptide. Nevertheless, the side chain interactions of the C-terminal tail with the DLG1 GK domain are similar to those of the phosphorylated LGN peptide.
Organizational Affiliation:
Department of Biophysics and Biochemistry, Graduate School of Science, The University of Tokyo, Tokyo 113-0033, Japan.