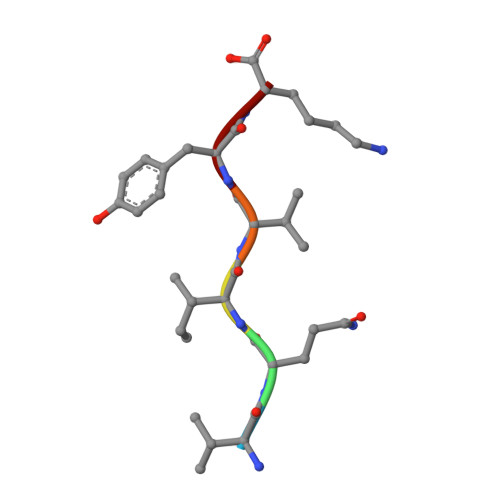
Towards a pharmacophore for amyloid.
Landau, M., Sawaya, M.R., Faull, K.F., Laganowsky, A., Jiang, L., Sievers, S.A., Liu, J., Barrio, J.R., Eisenberg, D.(2011) PLoS Biol 9: e1001080-e1001080
- PubMed: 21695112
- DOI: https://doi.org/10.1371/journal.pbio.1001080
- Primary Citation of Related Structures:
3OVJ, 3OVL - PubMed Abstract:
Diagnosing and treating Alzheimer's and other diseases associated with amyloid fibers remains a great challenge despite intensive research. To aid in this effort, we present atomic structures of fiber-forming segments of proteins involved in Alzheimer's disease in complex with small molecule binders, determined by X-ray microcrystallography. The fiber-like complexes consist of pairs of β-sheets, with small molecules binding between the sheets, roughly parallel to the fiber axis. The structures suggest that apolar molecules drift along the fiber, consistent with the observation of nonspecific binding to a variety of amyloid proteins. In contrast, negatively charged orange-G binds specifically to lysine side chains of adjacent sheets. These structures provide molecular frameworks for the design of diagnostics and drugs for protein aggregation diseases.
Organizational Affiliation:
Howard Hughes Medical Institute, UCLA-DOE Institute for Genomics and Proteomics, Department of Biological Chemistry, University of California, Los Angeles, California, United States of America.