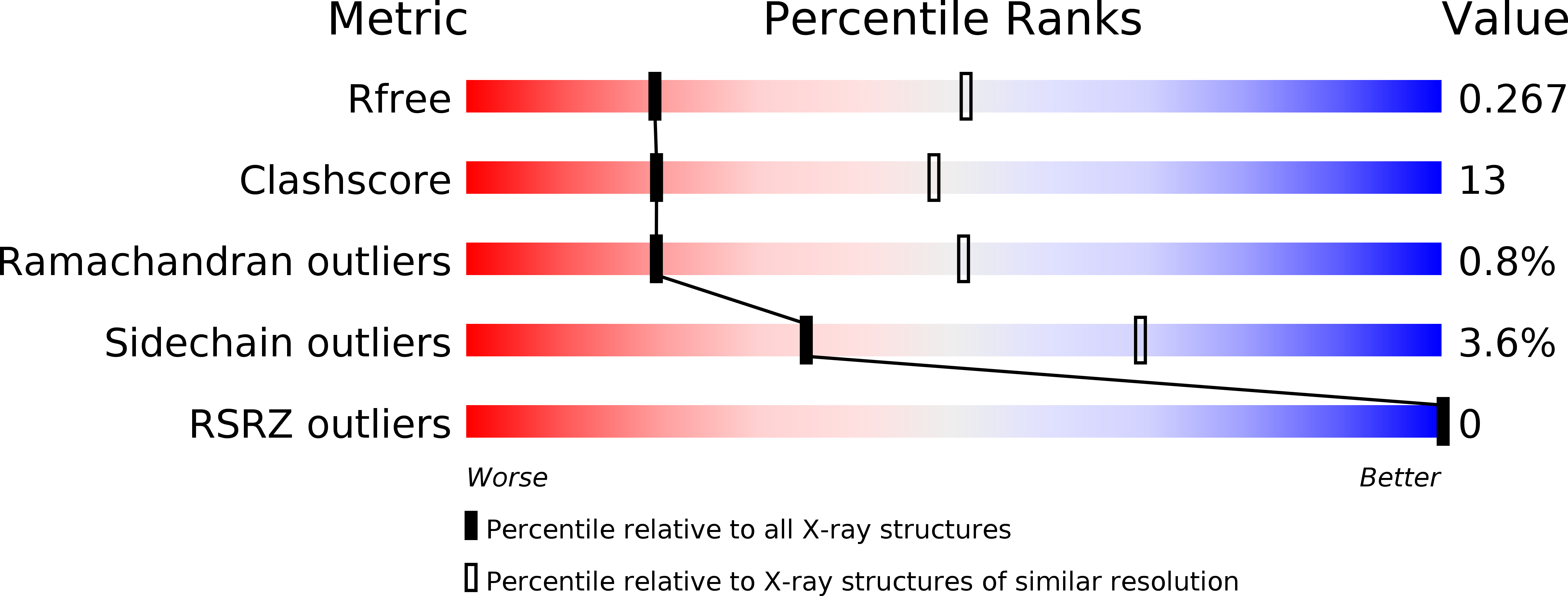
Co-Crystal Structures of PKG Ibeta (92-227) with cGMP and cAMP Reveal the Molecular Details of Cyclic-Nucleotide Binding
Kim, J.J., Casteel, D.E., Huang, G., Kwon, T.H., Ren, R.K., Zwart, P., Headd, J.J., Brown, N.G., Chow, D.C., Palzkill, T., Kim, C.(2011) PLoS One 6: e18413-e18413
- PubMed: 21526164
- DOI: https://doi.org/10.1371/journal.pone.0018413
- Primary Citation of Related Structures:
3OCP, 3OD0, 3OGJ - PubMed Abstract:
Cyclic GMP-dependent protein kinases (PKGs) are central mediators of the NO-cGMP signaling pathway and phosphorylate downstream substrates that are crucial for regulating smooth muscle tone, platelet activation, nociception and memory formation. As one of the main receptors for cGMP, PKGs mediate most of the effects of cGMP elevating drugs, such as nitric oxide-releasing agents and phosphodiesterase inhibitors which are used for the treatment of angina pectoris and erectile dysfunction, respectively.
Organizational Affiliation:
Department of Pharmacology, Baylor College of Medicine, Houston, Texas, United States of America.