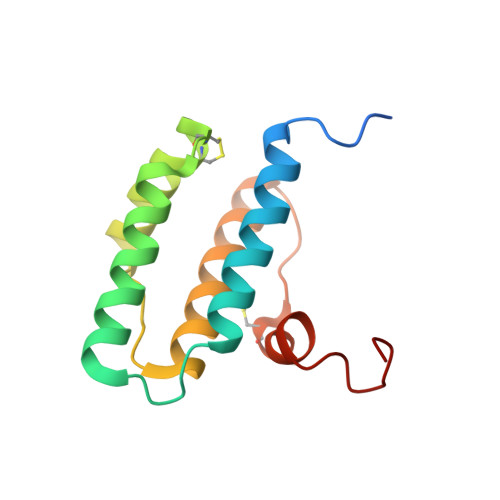
Molecular recognition and substrate mimicry drive the electron-transfer process between MIA40 and ALR.
Banci, L., Bertini, I., Calderone, V., Cefaro, C., Ciofi-Baffoni, S., Gallo, A., Kallergi, E., Lionaki, E., Pozidis, C., Tokatlidis, K.(2011) Proc Natl Acad Sci U S A 108: 4811-4816
- PubMed: 21383138
- DOI: https://doi.org/10.1073/pnas.1014542108
- Primary Citation of Related Structures:
3O55 - PubMed Abstract:
Oxidative protein folding in the mitochondrial intermembrane space requires the transfer of a disulfide bond from MIA40 to the substrate. During this process MIA40 is reduced and regenerated to a functional state through the interaction with the flavin-dependent sulfhydryl oxidase ALR. Here we present the mechanistic basis of ALR-MIA40 interaction at atomic resolution by biochemical and structural analyses of the mitochondrial ALR isoform and its covalent mixed disulfide intermediate with MIA40. This ALR isoform contains a folded FAD-binding domain at the C-terminus and an unstructured, flexible N-terminal domain, weakly and transiently interacting one with the other. A specific region of the N-terminal domain guides the interaction with the MIA40 substrate binding cleft (mimicking the interaction of the substrate itself), without being involved in the import of ALR. The hydrophobicity-driven binding of this region ensures precise protein-protein recognition needed for an efficient electron transfer process.
Organizational Affiliation:
Magnetic Resonance Center (CERM), University of Florence, Via Luigi Sacconi 6, Sesto Fiorentino, 50019 Florence, Italy. banci@cerm.unifi.it