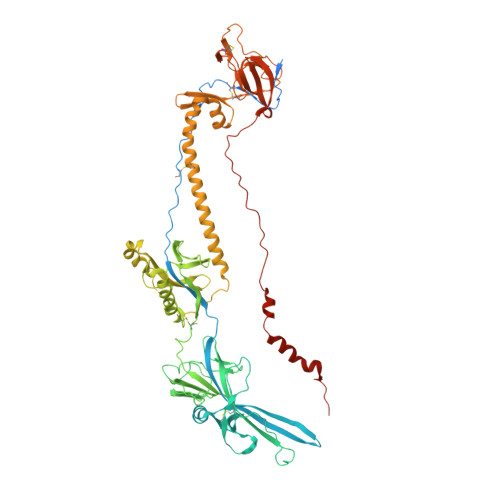
Structural basis of local, pH-dependent conformational changes in glycoprotein B from herpes simplex virus type 1.
Stampfer, S.D., Lou, H., Cohen, G.H., Eisenberg, R.J., Heldwein, E.E.(2010) J Virol 84: 12924-12933
- PubMed: 20943984
- DOI: https://doi.org/10.1128/JVI.01750-10
- Primary Citation of Related Structures:
3NW8, 3NWA, 3NWD, 3NWF - PubMed Abstract:
Herpesviruses enter cells by membrane fusion either at the plasma membrane or in endosomes, depending on the cell type. Glycoprotein B (gB) is a conserved component of the multiprotein herpesvirus fusion machinery and functions as a fusion protein, with two internal fusion loops, FL1 and FL2. We determined the crystal structures of the ectodomains of two FL1 mutants of herpes simplex virus type 1 (HSV-1) gB to clarify whether their fusion-null phenotypes were due to global or local effects of the mutations on the structure of the gB ectodomain. Each mutant has a single point mutation of a hydrophobic residue in FL1 that eliminates the hydrophobic side chain. We found that neither mutation affected the conformation of FL1, although one mutation slightly altered the conformation of FL2, and we conclude that the fusion-null phenotype is due to the absence of a hydrophobic side chain at the mutated position. Because the ectodomains of the wild-type and the mutant forms of gB crystallized at both low and neutral pH, we were able to determine the effect of pH on gB conformation at the atomic level. For viruses that enter cells by endocytosis, the low pH of the endosome effects major conformational changes in their fusion proteins, thereby promoting fusion of the viral envelope with the endosomal membrane. We show here that upon exposure of gB to low pH, FL2 undergoes a major relocation, probably driven by protonation of a key histidine residue. Relocation of FL2, as well as additional small conformational changes in the gB ectodomain, helps explain previously noted changes in its antigenic and biochemical properties. However, no global pH-dependent changes in gB structure were detected in either the wild-type or the mutant forms of gB. Thus, low pH causes local conformational changes in gB that are very different from the large-scale fusogenic conformational changes in other viral fusion proteins. We propose that these conformational changes, albeit modest, play an important functional role during endocytic entry of HSV.
Organizational Affiliation:
Department of Molecular Biology and Microbiology, Sackler School of Graduate Biomedical Sciences, Tufts University School of Medicine, 136 Harrison Avenue, Boston, MA 02111, USA.