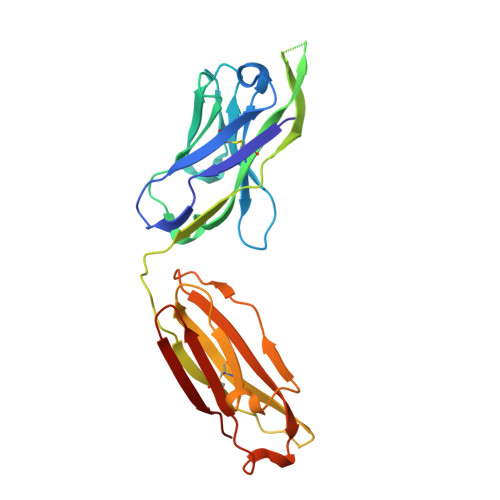
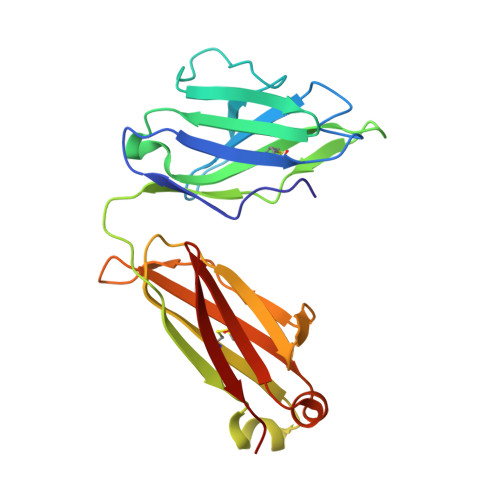
Crystal structure of PG16 and chimeric dissection with somatically related PG9: structure-function analysis of two quaternary-specific antibodies that effectively neutralize HIV-1.
Pancera, M., McLellan, J.S., Wu, X., Zhu, J., Changela, A., Schmidt, S.D., Yang, Y., Zhou, T., Phogat, S., Mascola, J.R., Kwong, P.D.(2010) J Virol 84: 8098-8110
- PubMed: 20538861
- DOI: https://doi.org/10.1128/JVI.00966-10
- Primary Citation of Related Structures:
3LRS, 3MME - PubMed Abstract:
HIV-1 resists neutralization by most antibodies. Two somatically related human antibodies, PG9 and PG16, however, each neutralize 70 to 80% of circulating HIV-1 isolates. Here we present the structure of the antigen-binding fragment of PG16 in monoclinic and orthorhombic lattices at 2.4 and 4.0 A, respectively, and use a combination of structural analysis, paratope dissection, and neutralization assessment to determine the functional relevance of three unusual PG9/PG16 features: N-linked glycosylation, extensive affinity maturation, and a heavy chain-third complementarity-determining region (CDR H3) that is one of the longest observed in human antibodies. Glycosylation extended off the side of the light chain variable domain and was not required for neutralization. The CDR H3 formed an axe-shaped subdomain, which comprised 42% of the CDR surface, with the axe head looming approximately 20 A above the other combining loops. Comprehensive sets of chimeric swaps between PG9 and PG16 of light chain, heavy chain, and CDR H3 were employed to decipher structure-function relationships. Chimeric swaps generally complemented functionally, with differences in PG9/PG16 neutralization related primarily to residue differences in CDR H3. Meanwhile, chimeric reversions to genomic V genes showed isolate-dependent effects, with affinity maturation playing a significant role in augmenting neutralization breadth (P = 0.036) and potency (P < 0.0001). The structural and functional details of extraordinary CDR H3 and extensive affinity maturation provide insights into the neutralization mechanism of and the elicitation pathway for broadly neutralizing antibodies like PG9 and PG16.
Organizational Affiliation:
Vaccine Research Center, National Institute of Allergy and Infectious Diseases, National Institutes of Health, Bethesda, Maryland 20892-3027, USA.