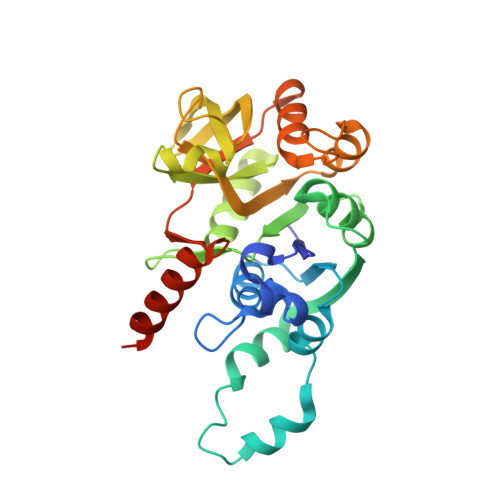
Structural basis for the reaction mechanism of UDP-glucose pyrophosphorylase
Kim, H., Choi, J., Kim, T., Lokanath, N.K., Ha, S.C., Suh, S.W., Hwang, H.-Y., Kim, K.K.(2010) Mol Cells 29: 397-405
- PubMed: 20238176
- DOI: https://doi.org/10.1007/s10059-010-0047-6
- Primary Citation of Related Structures:
3JUJ, 3JUK - PubMed Abstract:
UDP-glucose pyrophosphorylases (UGPase; EC 2.7.7.9) catalyze the conversion of UTP and glucose-1-phosphate to UDP-glucose and pyrophosphate and vice versa. Prokaryotic UGPases are distinct from their eukaryotic counterparts and are considered appropriate targets for the development of novel antibacterial agents since their product, UDP-glucose, is indispensable for the biosynthesis of virulence factors such as lipopolysaccharides and capsular polysaccharides. In this study, the crystal structures of UGPase from Helicobacter pylori (HpUGPase) were determined in apo- and UDP-glucose/Mg(2+)-bound forms at 2.9 A and 2.3 A resolutions, respectively. HpUGPase is a homotetramer and its active site is located in a deep pocket of each subunit. Magnesium ion is coordinated by Asp130, two oxygen atoms of phosphoryl groups, and three water molecules with octahedral geometry. Isothermal titration calorimetry analyses demonstrated that Mg(2+) ion plays a key role in the enzymatic activity of UGPase by enhancing the binding of UGPase to UTP or UDP-glucose, suggesting that this reaction is catalyzed by an ordered sequential Bi Bi mechanism. Furthermore, the crystal structure explains the specificity for uracil bases. The current structural study combined with functional analyses provides essential information for understanding the reaction mechanism of bacterial UGPases, as well as a platform for the development of novel antibacterial agents.
Organizational Affiliation:
Department of Molecular Cell Biology, Samsung Biomedical Research Institute, Sungkyunkwan University School of Medicine, Suwon, 440-746, Korea. kkim@med.skku.ac.kr