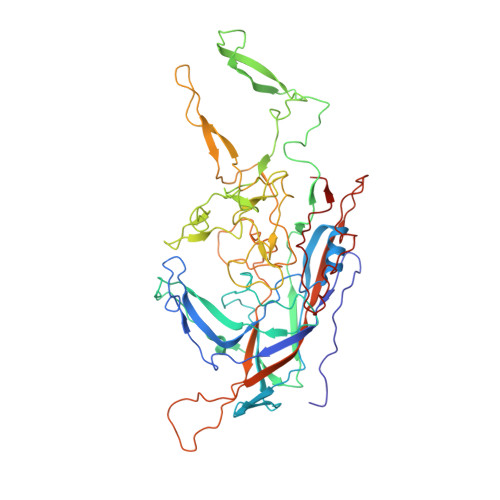
Electron microscopy analysis of a disaccharide analog complex reveals receptor interactions of adeno-associated virus.
Xie, Q., Spilman, M., Meyer, N.L., Lerch, T.F., Stagg, S.M., Chapman, M.S.(2013) J Struct Biol 184: 129-135
- PubMed: 24036405
- DOI: https://doi.org/10.1016/j.jsb.2013.09.004
- Primary Citation of Related Structures:
3J4P - PubMed Abstract:
Mechanistic studies of macromolecular complexes often feature X-ray structures of complexes with bound ligands. The attachment of adeno-associated virus (AAV) to cell surface glycosaminoglycans (GAGs) is an example that has not proven amenable to crystallography, because the binding of GAG analogs disrupts lattice contacts. The interactions of AAV with GAGs are of interest in mediating the cell specificity of AAV-based gene therapy vectors. Previous electron microscopy led to differing conclusions on the exact binding site and the existence of large ligand-induced conformational changes in the virus. Conformational changes are expected during cell entry, but it has remained unclear whether the electron microscopy provided evidence of their induction by GAG-binding. Taking advantage of automated data collection, careful processing and new methods of structure refinement, the structure of AAV-DJ complexed with sucrose octasulfate is determined by electron microscopy difference map analysis to 4.8Å resolution. At this higher resolution, individual sulfate groups are discernible, providing a stereochemical validation of map interpretation, and highlighting interactions with two surface arginines that have been implicated in genetic studies. Conformational changes induced by the SOS are modest and limited to the loop most directly interacting with the ligand. While the resolution attainable will depend on sample order and other factors, there are an increasing number of macromolecular complexes that can be studied by cryo-electron microscopy at resolutions beyond 5Å, for which the approaches used here could be used to characterize the binding of inhibitors and other small molecule effectors when crystallography is not tractable.
Organizational Affiliation:
Department of Biochemistry & Molecular Biology, School of Medicine, Oregon Health &v Science University, Portland, OR 97239-3098, USA.