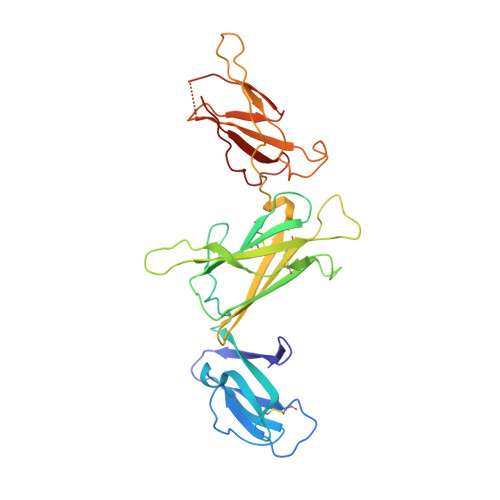
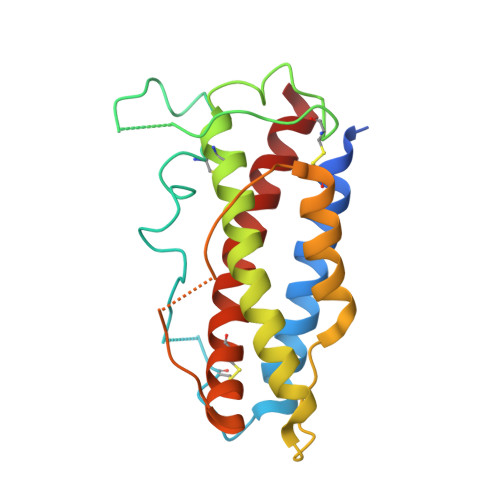
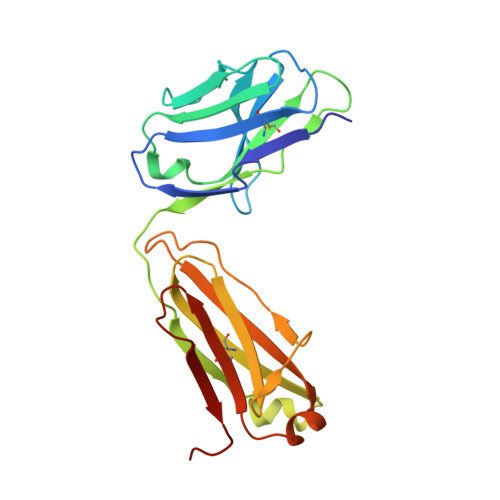
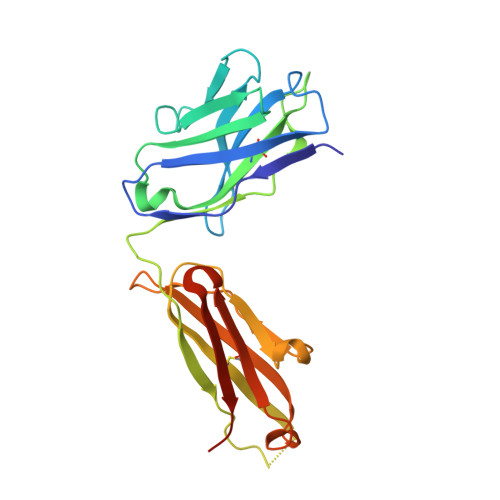
Structural basis for the dual recognition of IL-12 and IL-23 by ustekinumab.
Luo, J., Wu, S.J., Lacy, E.R., Orlovsky, Y., Baker, A., Teplyakov, A., Obmolova, G., Heavner, G.A., Richter, H.T., Benson, J.(2010) J Mol Biol 402: 797-812
- PubMed: 20691190
- DOI: https://doi.org/10.1016/j.jmb.2010.07.046
- Primary Citation of Related Structures:
3HMW, 3HMX - PubMed Abstract:
Interleukin (IL)-12 and IL-23 are heterodimeric proinflammatory cytokines that share a common p40 subunit, paired with p35 and p19 subunits, respectively. They represent an attractive class of therapeutic targets for the treatment of psoriasis and other immune-mediated diseases. Ustekinumab is a fully human monoclonal antibody (mAb) that binds specifically to IL-12/IL-23p40 and neutralizes human IL-12 and IL-23 bioactivity. The crystal structure of ustekinumab Fab (antigen binding fragment of mAb), in complex with human IL-12, has been determined by X-ray crystallography at 3.0 Å resolution. Ustekinumab Fab binds the D1 domain of the p40 subunit in a 1:1 ratio in the crystal, consistent with a 2 cytokines:1 mAb stoichiometry, as measured by isothermal titration calorimetry. The structure indicates that ustekinumab binds to the same epitope on p40 in both IL-12 and IL-23 with identical interactions. Mutational analyses confirm that several residues identified in the IL-12/IL-23p40 epitope provide important molecular binding interactions with ustekinumab. The electrostatic complementarity between the mAb antigen binding site and the p40 D1 domain epitope appears to play a key role in antibody/antigen recognition specificity. Interestingly, this structure also reveals significant structural differences in the p35 subunit and p35/p40 interface, compared with the published crystal structure of human IL-12, suggesting unusual and potentially functionally relevant structural flexibility of p35, as well as p40/p35 recognition. Collectively, these data describe unique observations about IL-12p35 and ustekinumab interactions with p40 that account for its dual binding and neutralization of IL-12 and IL-23.
Organizational Affiliation:
Centocor Research and Development, Inc., 145 King of Prussia Road, Radnor, PA 19087, USA. jluo@its.jnj.com