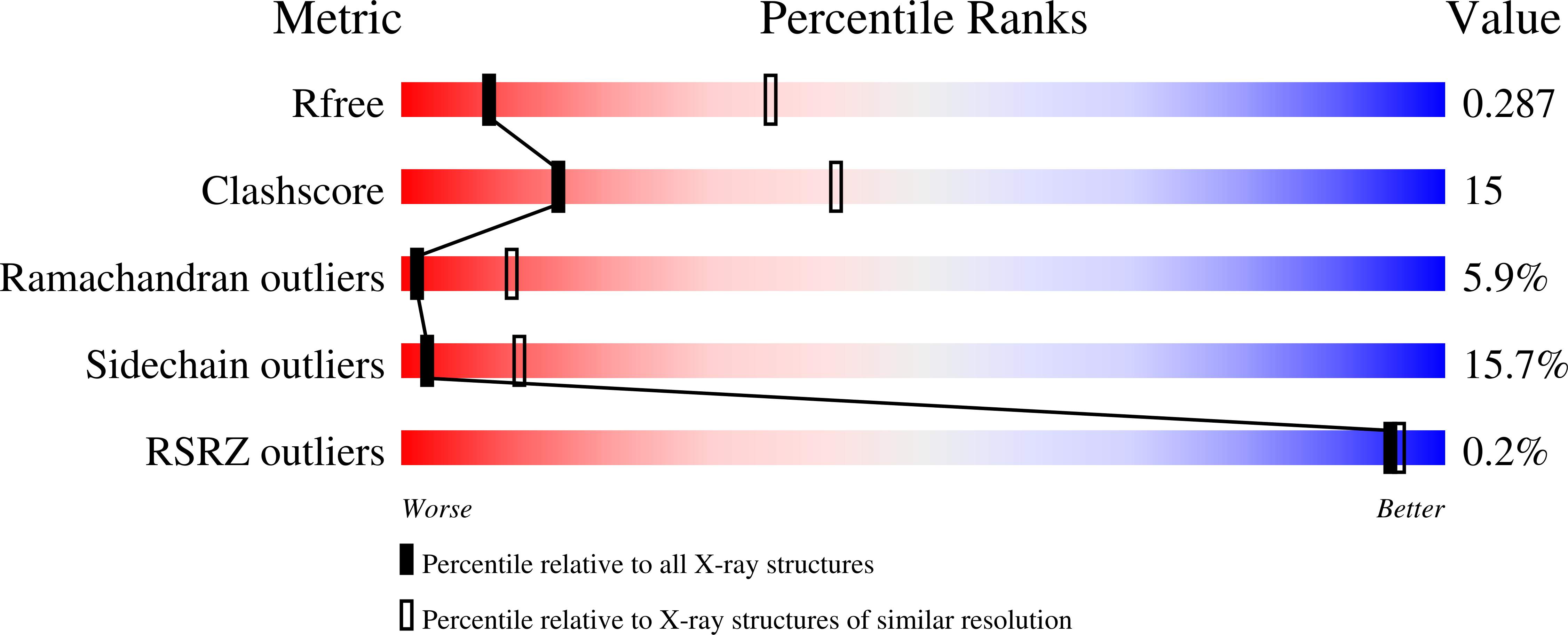
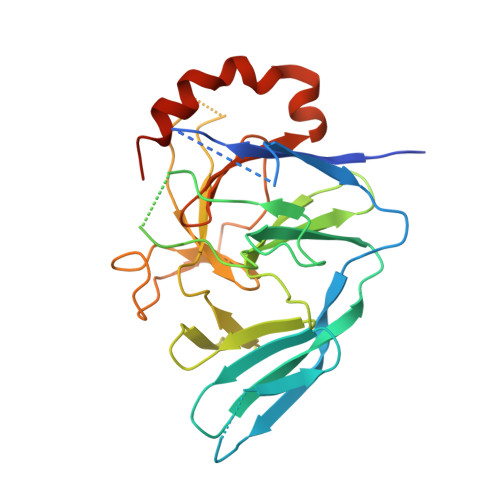
Pellino proteins contain a cryptic FHA domain that mediates interaction with phosphorylated IRAK1.
Lin, C.C., Huoh, Y.S., Schmitz, K.R., Jensen, L.E., Ferguson, K.M.(2008) Structure 16: 1806-1816
- PubMed: 19081057
- DOI: https://doi.org/10.1016/j.str.2008.09.011
- Primary Citation of Related Structures:
3EGA, 3EGB - PubMed Abstract:
Pellino proteins are RING E3 ubiquitin ligases involved in signaling events downstream of the Toll and interleukin-1 (IL-1) receptors, key initiators of innate immune and inflammatory responses. Pellino proteins associate with and ubiquitinate proteins in these pathways, including the interleukin-1 receptor associated kinase-1 (IRAK1). We determined the X-ray crystal structure of a Pellino2 fragment lacking only the RING domain. This structure reveals that the IRAK1-binding region of Pellino proteins consists largely of a previously unidentified forkhead-associated (FHA) domain. FHA domains are well-characterized phosphothreonine-binding modules, and this cryptic example in Pellino2 can drive interaction of this protein with phosphorylated IRAK1. The Pellino FHA domain is decorated with an unusual appendage or "wing" composed of two long inserts that lie within the FHA homology region. Delineating how this E3 ligase associates with substrates, and how these interactions are regulated by phosphorylation, is crucial for a complete understanding of Toll/IL-1 receptor signaling.
Organizational Affiliation:
Department of Physiology, University of Pennsylvania School of Medicine, Philadelphia, PA 19104, USA.