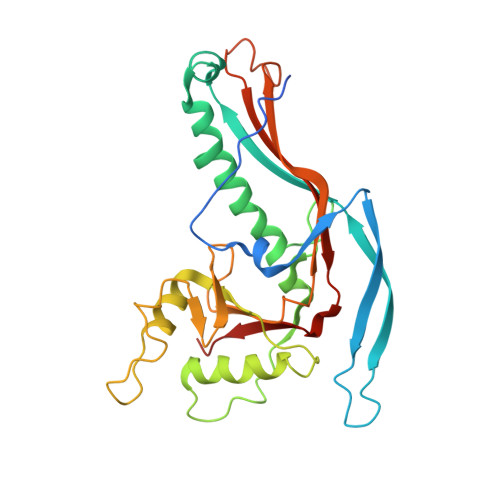
An unexpected twist in viral capsid maturation.
Gertsman, I., Gan, L., Guttman, M., Lee, K., Speir, J.A., Duda, R.L., Hendrix, R.W., Komives, E.A., Johnson, J.E.(2009) Nature 458: 646-650
- PubMed: 19204733
- DOI: https://doi.org/10.1038/nature07686
- Primary Citation of Related Structures:
3E8K - PubMed Abstract:
Lambda-like double-stranded (ds) DNA bacteriophage undergo massive conformational changes in their capsid shell during the packaging of their viral genomes. Capsid shells are complex organizations of hundreds of protein subunits that assemble into intricate quaternary complexes that ultimately are able to withstand over 50 atm of pressure during genome packaging. The extensive integration between subunits in capsids requires the formation of an intermediate complex, termed a procapsid, from which individual subunits can undergo the necessary refolding and structural rearrangements needed to transition to the more stable capsid. Although various mature capsids have been characterized at atomic resolution, no such procapsid structure is available for a dsDNA virus or bacteriophage. Here we present a procapsid X-ray structure at 3.65 A resolution, termed prohead II, of the lambda-like bacteriophage HK97, the mature capsid structure of which was previously solved to 3.44 A (ref. 2). A comparison of the two largely different capsid forms has unveiled an unprecedented expansion mechanism that describes the transition. Crystallographic and hydrogen/deuterium exchange data presented here demonstrate that the subunit tertiary structures are significantly different between the two states, with twisting and bending motions occurring in both helical and beta-sheet regions. We also identified subunit interactions at each three-fold axis of the capsid that are maintained throughout maturation. The interactions sustain capsid integrity during subunit refolding and provide a fixed hinge from which subunits undergo rotational and translational motions during maturation. Previously published calorimetric data of a closely related bacteriophage, P22, showed that capsid maturation was an exothermic process that resulted in a release of 90 kJ mol(-1) of energy. We propose that the major tertiary changes presented in this study reveal a structural basis for an exothermic maturation process probably present in many dsDNA bacteriophage and possibly viruses such as herpesvirus, which share the HK97 subunit fold.
Organizational Affiliation:
Department of Molecular Biology, The Scripps Research Institute, La Jolla, California 92037, USA.