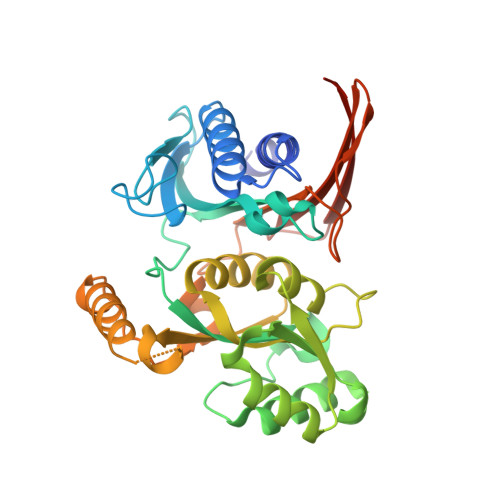
Structural and Biochemical Characterization of the Type II Fructose-1,6-bisphosphatase GlpX from Escherichia coli.
Brown, G., Singer, A., Lunin, V.V., Proudfoot, M., Skarina, T., Flick, R., Kochinyan, S., Sanishvili, R., Joachimiak, A., Edwards, A.M., Savchenko, A., Yakunin, A.F.(2009) J Biol Chem 284: 3784-3792
- PubMed: 19073594
- DOI: https://doi.org/10.1074/jbc.M808186200
- Primary Citation of Related Structures:
1NI9, 3BIG, 3BIH, 3D1R - PubMed Abstract:
Gluconeogenesis is an important metabolic pathway, which produces glucose from noncarbohydrate precursors such as organic acids, fatty acids, amino acids, or glycerol. Fructose-1,6-bisphosphatase, a key enzyme of gluconeogenesis, is found in all organisms, and five different classes of these enzymes have been identified. Here we demonstrate that Escherichia coli has two class II fructose-1,6-bisphosphatases, GlpX and YggF, which show different catalytic properties. We present the first crystal structure of a class II fructose-1,6-bisphosphatase (GlpX) determined in a free state and in the complex with a substrate (fructose 1,6-bisphosphate) or inhibitor (phosphate). The crystal structure of the ligand-free GlpX revealed a compact, globular shape with two alpha/beta-sandwich domains. The core fold of GlpX is structurally similar to that of Li+-sensitive phosphatases implying that they have a common evolutionary origin and catalytic mechanism. The structure of the GlpX complex with fructose 1,6-bisphosphate revealed that the active site is located between two domains and accommodates several conserved residues coordinating two metal ions and the substrate. The third metal ion is bound to phosphate 6 of the substrate. Inorganic phosphate strongly inhibited activity of both GlpX and YggF, and the crystal structure of the GlpX complex with phosphate demonstrated that the inhibitor molecule binds to the active site. Alanine replacement mutagenesis of GlpX identified 12 conserved residues important for activity and suggested that Thr(90) is the primary catalytic residue. Our data provide insight into the molecular mechanisms of the substrate specificity and catalysis of GlpX and other class II fructose-1,6-bisphosphatases.
Organizational Affiliation:
Banting and Best Department of Medical Research, University of Toronto, Toronto, Ontario M5G 1L6, Canada.