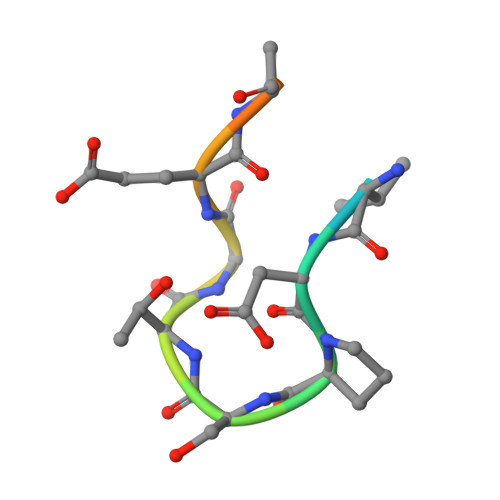
The selective autophagy substrate p62 activates the stress responsive transcription factor Nrf2 through inactivation of Keap1
Komatsu, M., Kurokawa, H., Waguri, S., Taguchi, K., Kobayashi, A., Ichimura, Y., Sou, Y.S., Ueno, I., Sakamoto, A., Tong, K.I., Kim, M., Nishito, Y., Iemura, S., Natsume, T., Ueno, T., Kominami, E., Motohashi, H., Tanaka, K., Yamamoto, M.(2010) Nat Cell Biol 12: 213-223
- PubMed: 20173742
- DOI: https://doi.org/10.1038/ncb2021
- Primary Citation of Related Structures:
3ADE - PubMed Abstract:
Impaired selective turnover of p62 by autophagy causes severe liver injury accompanied by the formation of p62-positive inclusions and upregulation of detoxifying enzymes. These phenotypes correspond closely to the pathological conditions seen in human liver diseases, including alcoholic hepatitis and hepatocellular carcinoma. However, the molecular mechanisms and pathophysiological processes in these events are still unknown. Here we report the identification of a novel regulatory mechanism by p62 of the transcription factor Nrf2, whose target genes include antioxidant proteins and detoxification enzymes. p62 interacts with the Nrf2-binding site on Keap1, a component of Cullin-3-type ubiquitin ligase for Nrf2. Thus, an overproduction of p62 or a deficiency in autophagy competes with the interaction between Nrf2 and Keap1, resulting in stabilization of Nrf2 and transcriptional activation of Nrf2 target genes. Our findings indicate that the pathological process associated with p62 accumulation results in hyperactivation of Nrf2 and delineates unexpected roles of selective autophagy in controlling the transcription of cellular defence enzyme genes.
- Laboratory of Frontier Science, Tokyo Metropolitan Institute of Medical Science, Bunkyo-ku, Tokyo 113-8613, Japan. komatsu-ms@igakuken.or.jp
Organizational Affiliation: