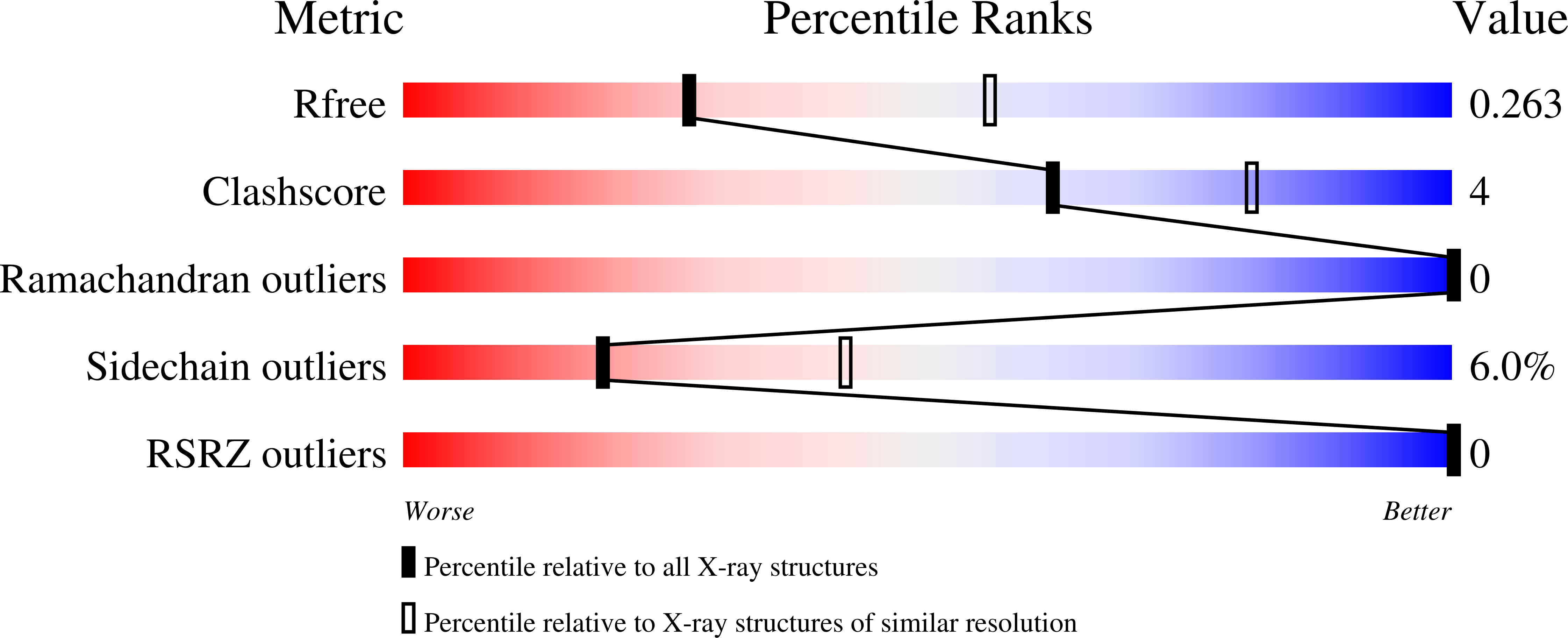
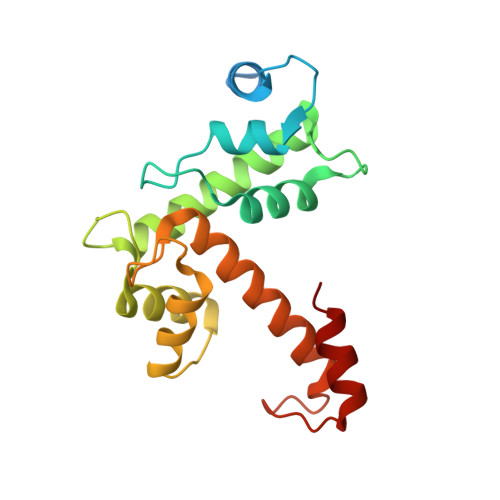
Structural Basis for Ca(2+)-Dependent Formation of ALG-2/Alix Peptide Complex: Ca(2+)/EF3-Driven Arginine Switch Mechanism
Suzuki, H., Kawasaki, M., Inuzuka, T., Okumura, M., Kakiuchi, T., Shibata, H., Wakatsuki, S., Maki, M.(2008) Structure 16: 1562-1573
- PubMed: 18940611
- DOI: https://doi.org/10.1016/j.str.2008.07.012
- Primary Citation of Related Structures:
2ZN8, 2ZN9, 2ZND, 2ZNE - PubMed Abstract:
ALG-2 belongs to the penta-EF-hand (PEF) protein family and interacts with various intracellular proteins, such as Alix and TSG101, that are involved in endosomal sorting and HIV budding. Through X-ray crystallography, we solved the structures of Ca(2+)-free and -bound forms of N-terminally truncated human ALG-2 (des3-20ALG-2), Zn(2+)-bound form of full-length ALG-2, and the structure of the complex between des3-23ALG-2 and the peptide corresponding to Alix799-814 in Zn(2+)-bound form. Binding of Ca(2+) to EF3 enables the side chain of Arg125, present in the loop connecting EF3 and EF4, to move enough to make a primary hydrophobic pocket accessible to the critical PPYP motif, which partially overlaps with the GPP motif for the binding of Cep55 (centrosome protein 55 kDa). Based on these results, together with the results of in vitro binding assay with mutant ALG-2 and Alix proteins, we propose a Ca(2+)/EF3-driven arginine switch mechanism for ALG-2 binding to Alix.
Organizational Affiliation:
Department of Applied Molecular Biosciences, Graduate School of Bioagricultural Sciences, Nagoya University, Nagoya 464-8601, Japan.