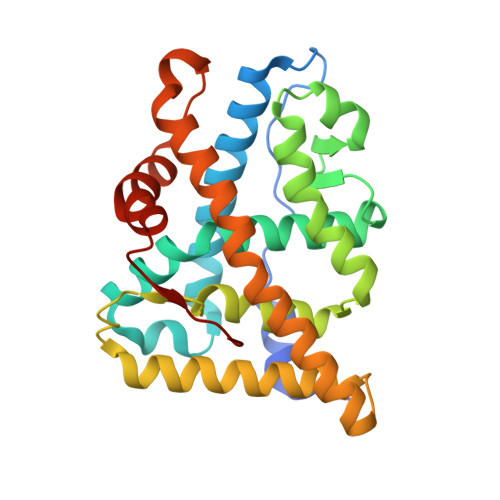
The X-Ray Structure of Ru486 Bound to the Progesterone Receptor in a Destabilized Agonistic Conformation.
Raaijmakers, H.C.A., Versteegh, J., Uitdehaag, J.C.M.(2009) J Biol Chem 284: 19572
- PubMed: 19372222
- DOI: https://doi.org/10.1074/jbc.M109.007872
- Primary Citation of Related Structures:
2W8Y - PubMed Abstract:
Here we describe the 1.95 A structure of the clinically used antiprogestin RU486 (mifepristone) in complex with the progesterone receptor (PR). The structure was obtained by taking a crystal of the PR ligand binding domain containing the agonist norethindrone and soaking it in a solution containing the antagonist RU486 for extended times. Clear ligand exchange could be observed in one copy of the PR ligand binding domain dimer in the crystal. RU486 binds while PR is in an agonistic conformation without displacing helix 12. Although this is probably because of the constraints of the crystal lattice, it demonstrates that helix 12 displacement is not a prerequisite for RU486 binding. Interestingly, B-factor analysis clearly shows that helix 12 becomes more flexible after RU486 binding, suggesting that RU486, being a model antagonist, does not induce one fixed conformation of helix 12 but changes its positional equilibrium. This conclusion is confirmed by comparing the structures of RU486 bound to PR and RU486 bound to the glucocorticoid receptor.
Organizational Affiliation:
Departments of Molecular Design and Informatics, Molecular Pharmacology and DMPK, Schering-Plough Research Institute, P. O. Box 20, 5340 BH Oss, The Netherlands.