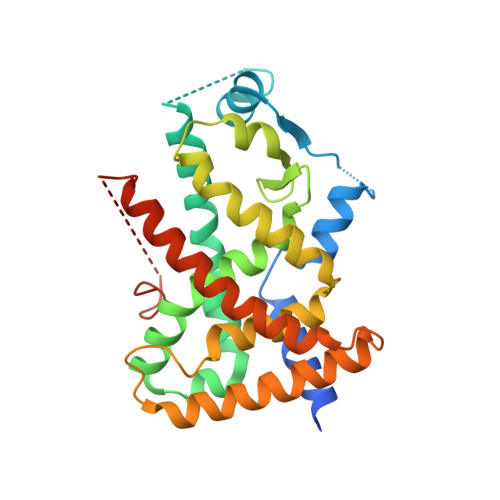
Design of a partial PPARdelta agonist.
Pettersson, I., Ebdrup, S., Havranek, M., Pihera, P., Korinek, M., Mogensen, J.P., Jeppesen, C.B., Johansson, E., Sauerberg, P.(2007) Bioorg Med Chem Lett 17: 4625-4629
- PubMed: 17560785
- DOI: https://doi.org/10.1016/j.bmcl.2007.05.079
- Primary Citation of Related Structures:
2Q5G - PubMed Abstract:
Structure based ligand design was used in order to design a partial agonist for the PPARdelta receptor. The maximum activation in the transactivation assay was reduced from 87% to 39%. The crystal structure of the ligand binding domain of the PPARdelta receptor in complex with compound 2 was determined in order to understand the structural changes which gave rise to the decrease in maximum activation.
Organizational Affiliation:
Novo Nordisk A/S, Novo Nordisk Park, 2760 Måløv, Denmark. inpe@novonordisk.com