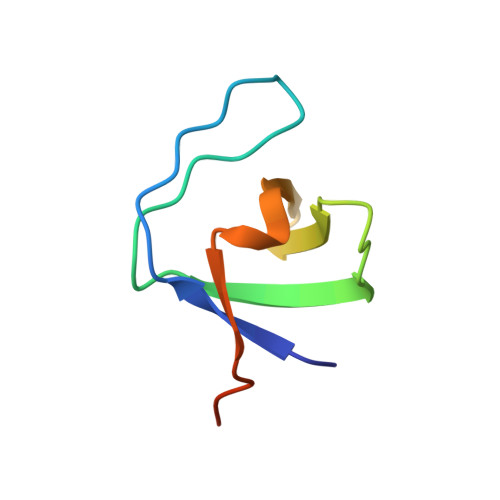
Solution structure of the first SH3 domain of human vinexin and its interaction with vinculin peptides
Zhang, J., Li, X., Yao, B., Shen, W., Sun, H., Xu, C., Wu, J., Shi, Y.(2007) Biochem Biophys Res Commun 357: 931-937
- PubMed: 17467669
- DOI: https://doi.org/10.1016/j.bbrc.2007.04.029
- Primary Citation of Related Structures:
2NWM - PubMed Abstract:
Solution structure of the first Src homology (SH) 3 domain of human vinexin (V_SH3_1) was determined using nuclear magnetic resonance (NMR) method and revealed that it was a canonical SH3 domain, which has a typical beta-beta-beta-beta-alpha-beta fold. Using chemical shift perturbation and surface plasmon resonance experiments, we studied the binding properties of the SH3 domain with two different peptides from vinculin hinge regions: P856 and P868. The observations illustrated slightly different affinities of the two peptides binding to V_SH3_1. The interaction between P868 and V_SH3_1 belonged to intermediate exchange with a modest binding affinity, while the interaction between P856 and V_SH3_1 had a low binding affinity. The structure and ligand-binding interface of V_SH3_1 provide a structural basis for the further functional study of this important molecule.
Organizational Affiliation:
Hefei National Laboratory for Physical Sciences at Microscale and School of Life Sciences, University of Science and Technology of China, Hefei, Anhui 230026, People's Republic of China.