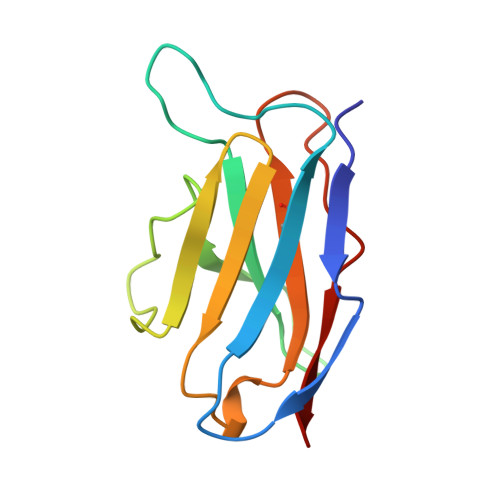
A domain flip as a result of a single amino-acid substitution.
Pokkuluri, P.R., Huang, D.B., Raffen, R., Cai, X., Johnson, G., Stevens, P.W., Stevens, F.J., Schiffer, M.(1998) Structure 6: 1067-1073
- PubMed: 9739086
- DOI: https://doi.org/10.1016/s0969-2126(98)00107-5
- Primary Citation of Related Structures:
2LVE, 3LVE, 4LVE - PubMed Abstract:
The self-assembly properties of beta domains are important features of diverse classes of proteins that include cell-adhesion molecules, surface receptors and the immunoglobulin superfamily. Immunoglobulin light-chain variable domains are well suited to the study of structural factors that determine dimerization, including how residues at the interface influence the preferred dimer arrangement.
Organizational Affiliation:
Center for Mechanistic Biology and Biotechnology, Argonne National Laboratory, IL 60439, USA.