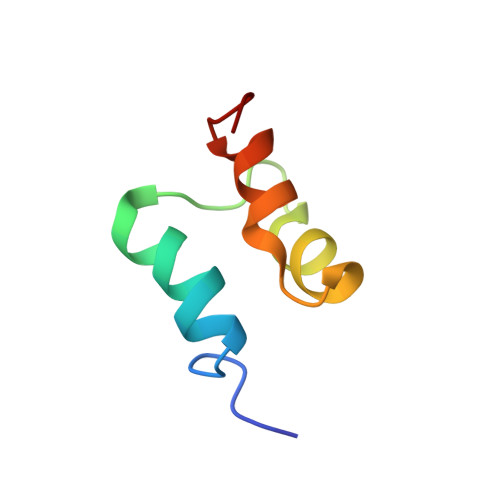
Conformation and dynamics of the three-helix bundle UBA domain of p62 from experiment and simulation.
Evans, C.L., Long, J.E., Gallagher, T.R., Hirst, J.D., Searle, M.S.(2007) Proteins 71: 227-240
- PubMed: 17932931
- DOI: https://doi.org/10.1002/prot.21692
- Primary Citation of Related Structures:
2K0B - PubMed Abstract:
The ubiquitin associated domain of p62 is a small three-helix bundle of approximately 50 residues that mediates the recognition of polyubiquitin chains and ubiquitylated substrates. The solution structure of a 52 residue construct containing this domain has been characterized using heteronuclear nuclear magnetic resonance (NMR) methods. The resulting ensemble of NMR-derived structures was used in molecular dynamics (MD) simulations to investigate the equilibrium conformation and dynamics of this domain. NOE and (15)N relaxation data have been used to validate the structural ensemble produced by the MD simulations and show a good correlation for residues in regions of secondary structure. A similar approach was taken using an ensemble of structures from the MD simulations to calculate electronic circular dichroism (CD) and IR spectra from first principles with an encouraging correlation with the experimental CD and IR data.
Organizational Affiliation:
School of Chemistry, University Park, University of Nottingham, Nottingham NG7 2RD, United Kingdom.