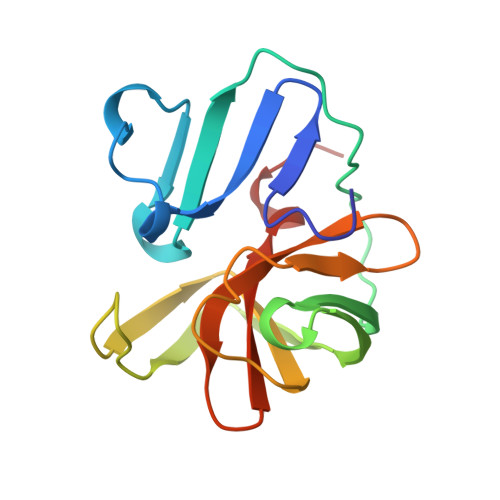
The structure of the 2A proteinase from a common cold virus: a proteinase responsible for the shut-off of host-cell protein synthesis.
Petersen, J.F., Cherney, M.M., Liebig, H.D., Skern, T., Kuechler, E., James, M.N.(1999) EMBO J 18: 5463-5475
- PubMed: 10523291
- DOI: https://doi.org/10.1093/emboj/18.20.5463
- Primary Citation of Related Structures:
2HRV - PubMed Abstract:
The crystal structure of the 2A proteinase from human rhinovirus serotype 2 (HRV2-2A(pro)) has been solved to 1.95 A resolution. The structure has an unusual, although chymotrypsin-related, fold comprising a unique four-stranded beta sheet as the N-terminal domain and a six-stranded beta barrel as the C-terminal domain. A tightly bound zinc ion, essential for the stability of HRV2-2A(pro), is tetrahedrally coordinated by three cysteine sulfurs and one histidine nitrogen. The active site consists of a catalytic triad formed by His18, Asp35 and Cys106. Asp35 is additionally involved in an extensive hydrogen-bonding network. Modelling studies reveal a substrate-induced fit that explains the specificity of the subsites S4, S2, S1 and S1'. The structure of HRV2-2A(pro) suggests the mechanism of the cis cleavage and its release from the polyprotein.
Organizational Affiliation:
MRC Group in Protein Structure and Function, Department of Biochemistry, University of Alberta, Edmonton, AB, Canada T6G 2H7.