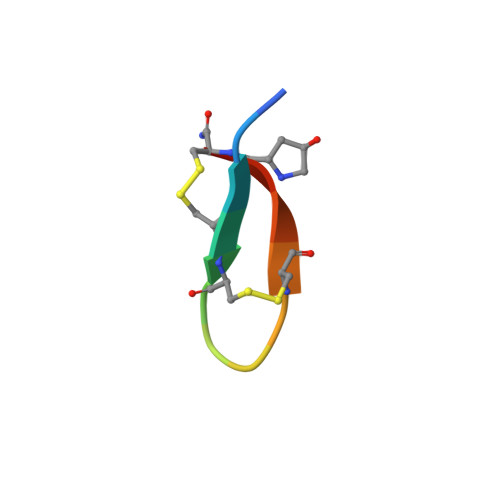
Solution structure of chi-conopeptide MrIA, a modulator of the human norepinephrine transporter
Nilsson, K.P., Lovelace, E.S., Caesar, C.E., Tynngard, N., Alewood, P.F., Johansson, H.M., Sharpe, I.A., Lewis, R.J., Daly, N.L., Craik, D.J.(2005) Biopolymers 80: 815-823
- PubMed: 15931669
- DOI: https://doi.org/10.1002/bip.20302
- Primary Citation of Related Structures:
2EW4 - PubMed Abstract:
The chi-conopeptides MrIA and MrIB are 13-residue peptides with two disulfide bonds that inhibit human and rat norepinephrine transporter systems and are of significant interest for the design of novel drugs involved in pain treatment. In the current study we have determined the solution structure of MrIA using NMR spectroscopy. The major element of secondary structure is a beta-hairpin with the two strands connected by an inverse gamma-turn. The residues primarily involved in activity have previously been shown to be located in the turn region (Sharpe, I. A.; Palant, E.; Schroder, C. I.; Kaye, D. M.; Adams, D. J.; Alewood, P. F.; Lewis, R. J. J Biol Chem 2003, 278, 40317-40323), which appears to be more flexible than the beta-strands based on disorder in the ensemble of calculated structures. Analogues of MrIA with N-terminal truncations indicate that the N-terminal residues play a role in defining a stable conformation and the native disulfide connectivity. In particular, noncovalent interactions between Val3 and Hyp12 are likely to be involved in maintaining a stable conformation. The N-terminus also affects activity, as a single N-terminal deletion introduced additional pharmacology at rat vas deferens, while deleting the first two amino acids reduced chi-conopeptide potency.
Organizational Affiliation:
Institute for Molecular Bioscience, University of Queensland, Brisbane 4072, Australia.