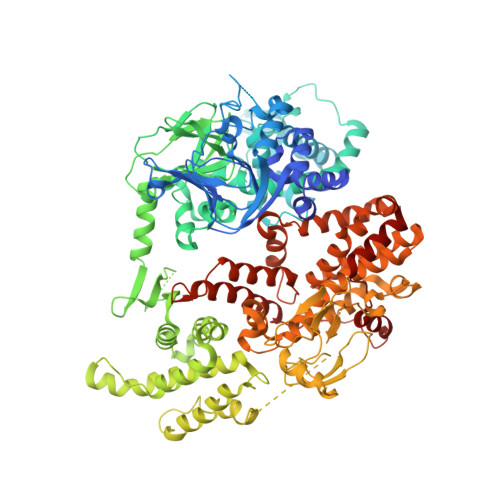
Structure of the Cyclic-AMP Responsive Exchange Factor Epac2 in its Auto-Inhibited State
Rehmann, H., Das, J., Knipscheer, P., Wittinghofer, A., Bos, J.L.(2006) Nature 439: 625
- PubMed: 16452984
- DOI: https://doi.org/10.1038/nature04468
- Primary Citation of Related Structures:
2BYV - PubMed Abstract:
Epac proteins (exchange proteins directly activated by cAMP) are guanine-nucleotide-exchange factors (GEFs) for the small GTP-binding proteins Rap1 and Rap2 that are directly regulated by the second messenger cyclic AMP and function in the control of diverse cellular processes, including cell adhesion and insulin secretion. Here we report the three-dimensional structure of full-length Epac2, a 110-kDa protein that contains an amino-terminal regulatory region with two cyclic-nucleotide-binding domains and a carboxy-terminal catalytic region. The structure was solved in the absence of cAMP and shows the auto-inhibited state of Epac. The regulatory region is positioned with respect to the catalytic region by a rigid, tripartite beta-sheet-like structure we refer to as the 'switchboard' and an ionic interaction we call the 'ionic latch'. As a consequence of this arrangement, the access of Rap to the catalytic site is sterically blocked. Mutational analysis suggests a model for cAMP-induced Epac activation with rigid body movement of the regulatory region, the features of which are universally conserved in cAMP-regulated proteins.
Organizational Affiliation:
Department of Physiological Chemistry and Centre for Biomedical Genetics, University Medical Center Utrecht, Universiteitsweg 100, 3584 CG Utrecht, The Netherlands.