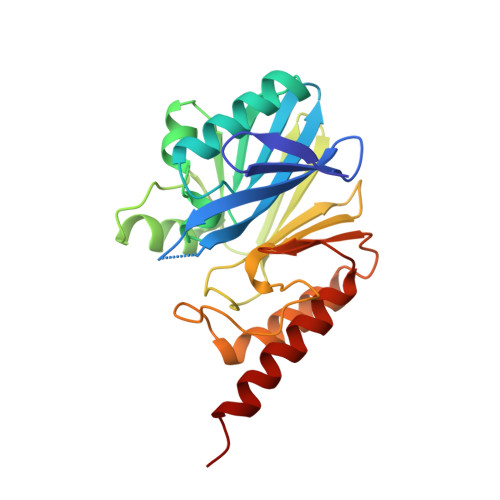
Crystal structure of the wide-spectrum binuclear zinc beta-lactamase from Bacteroides fragilis.
Concha, N.O., Rasmussen, B.A., Bush, K., Herzberg, O.(1996) Structure 4: 823-836
- PubMed: 8805566
- DOI: https://doi.org/10.1016/s0969-2126(96)00089-5
- Primary Citation of Related Structures:
1ZNB - PubMed Abstract:
The metallo-beta-lactamase from Bacteroides fragilis hydrolyzes a wide range of beta-lactam antibiotics, and is not clinically susceptible to any known beta-lactamase inhibitors. B. fragilis is associated with post-surgery hospital infections, and there has been a recent report of plasmid-mediated dissemination of the enzyme. Effective inhibitors are therefore urgently needed. Knowledge of the three-dimensional structure will aid in the drug design effort. The crystal structure of the enzyme has been determined by using multiwavelength anomalous diffraction at the zinc absorption edge and refined to 1.85 A resolution. The structure is a four-layer alpha/beta/beta/alpha molecule. The active site, found at the edge of the beta sandwich contains a binuclear zinc center with several novel features. One zinc is tetrahedrally coordinated, the other has a trigonal bipyramidal coordination; a water/hydroxide molecule serves as a ligand for both metals. The residues that coordinate the two zincs are invariant in all metallo-beta-lactamases that have been sequenced, except for two conservative replacements. Despite the existence of the pattern for binuclear zinc binding, the reported structure of the Bacillus cereus enzyme contains only a single zinc. Structural analysis indicates that affinity for the penta-coordinated zinc can be modulated by neighboring residues, perhaps explaining the absence of the second zinc in the B. cereus structure. Models of bound substrates suggest that the active-site channel can accommodate a wide variety of beta-lactams. We propose that the zinc cluster prepares an hydroxide, probably the hydroxide that ligates both zincs, for nucleophilic attack on the carbonyl carbon atom of the beta-lactam. The resulting negatively charged tetrahedral intermediate implicated in catalysis is stabilized by an oxyanion hole formed by the side chain of the invariant Asn 193 and the tetrahedral zinc.
Organizational Affiliation:
Center for Advanced Research in Biotechnology, University of Maryland Biotechnology Institute, Rockville 20850, USA.