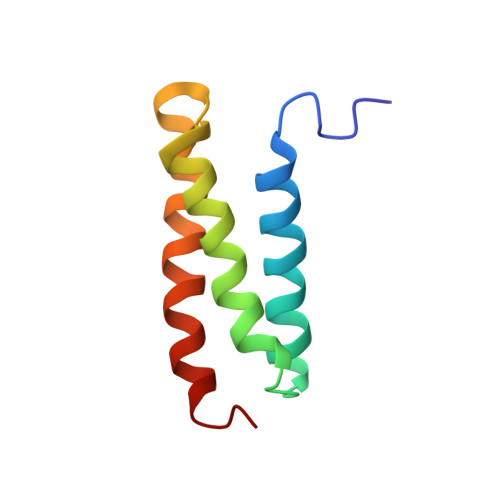
Structural characterization of the MIT domain from human Vps4b
Takasu, H., Jee, J.G., Ohno, A., Goda, N., Fujiwara, K., Tochio, H., Shirakawa, M., Hiroaki, H.(2005) Biochem Biophys Res Commun 334: 460-465
- PubMed: 16018968
- DOI: https://doi.org/10.1016/j.bbrc.2005.06.110
- Primary Citation of Related Structures:
1WR0 - PubMed Abstract:
The microtubule interacting and trafficking (MIT) domain is a small protein module of unknown function that is conserved in proteins of diverse function, such as Vps4, sorting nexin 15 (SNX15), and spastin. One non-synonymous single nucleotide polymorphism was reported, which results in a Ile58-to-Met (I58M) substitution in hVps4b. Here, we have determined the solution structure of the MIT domain isolated from the NH(2)-terminus of human Vps4b, an AAA-ATPase involved in multivesicular body formation. The MIT domain adopts an 'up-and-down' three-helix bundle. Comparison with the sequences of other MIT domains clearly shows that the residues involved in inter-helical contacts are well conserved. The Ile58-to-Met substitution resulted a substantial thermal instability. In addition, we found a shallow crevice between helices A and C that may serve as a protein-binding site. We propose that the MIT domain serves as a putative adaptor domain for the ESCRT-III complex involved in endosomal trafficking.
Organizational Affiliation:
International Graduate School of Arts and Sciences, Yokohama City University, Yokohama, Kanagawa 230 0045, Japan.