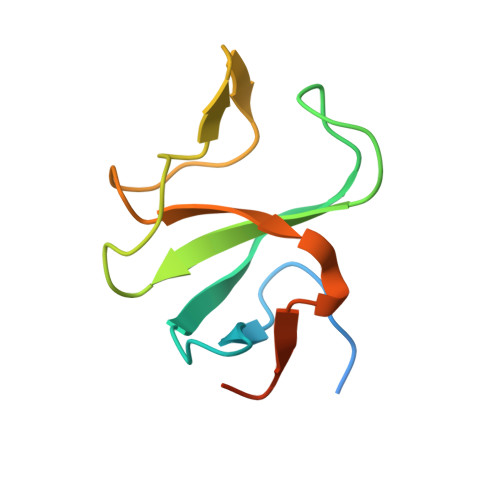
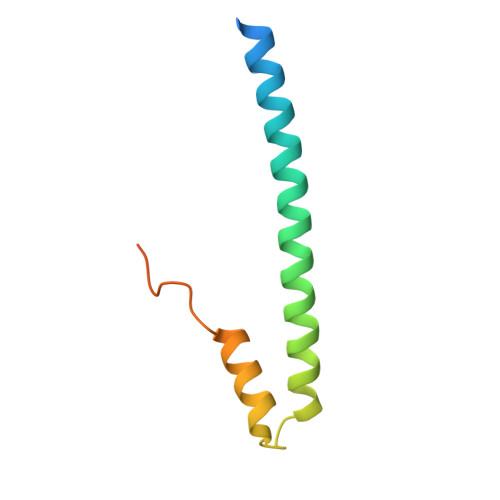
Structural Basis for the Activation of Microtubule Assembly by the EB1 and p150(Glued) Complex
Hayashi, I., Wilde, A., Mal, T.K., Ikura, M.(2005) Mol Cell 19: 449-460
- PubMed: 16109370
- DOI: https://doi.org/10.1016/j.molcel.2005.06.034
- Primary Citation of Related Structures:
1TXQ - PubMed Abstract:
Plus-end tracking proteins, such as EB1 and the dynein/dynactin complex, regulate microtubule dynamics. These proteins are thought to stabilize microtubules by forming a plus-end complex at microtubule growing ends with ill-defined mechanisms. Here we report the crystal structure of two plus-end complex components, the carboxy-terminal dimerization domain of EB1 and the microtubule binding (CAP-Gly) domain of the dynactin subunit p150Glued. Each molecule of the EB1 dimer contains two helices forming a conserved four-helix bundle, while also providing p150Glued binding sites in its flexible tail region. Combining crystallography, NMR, and mutational analyses, our studies reveal the critical interacting elements of both EB1 and p150Glued, whose mutation alters microtubule polymerization activity. Moreover, removal of the key flexible tail from EB1 activates microtubule assembly by EB1 alone, suggesting that the flexible tail negatively regulates EB1 activity. We, therefore, propose that EB1 possesses an auto-inhibited conformation, which is relieved by p150Glued as an allosteric activator.
Organizational Affiliation:
Division of Molecular and Structural Biology, Ontario Cancer Institute and Department of Medical Biophysics, University of Toronto, 610 University Avenue, Toronto, Ontario, M5G 2M9, Canada. ihayashi@uhnres.utoronto.ca