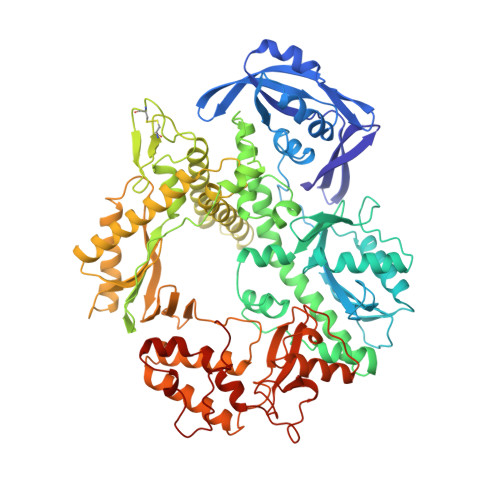
Crystal structure of a thermostable type B DNA polymerase from Thermococcus gorgonarius.
Hopfner, K.P., Eichinger, A., Engh, R.A., Laue, F., Ankenbauer, W., Huber, R., Angerer, B.(1999) Proc Natl Acad Sci U S A 96: 3600-3605
- PubMed: 10097083
- DOI: https://doi.org/10.1073/pnas.96.7.3600
- Primary Citation of Related Structures:
1TGO - PubMed Abstract:
Most known archaeal DNA polymerases belong to the type B family, which also includes the DNA replication polymerases of eukaryotes, but maintain high fidelity at extreme conditions. We describe here the 2.5 A resolution crystal structure of a DNA polymerase from the Archaea Thermococcus gorgonarius and identify structural features of the fold and the active site that are likely responsible for its thermostable function. Comparison with the mesophilic B type DNA polymerase gp43 of the bacteriophage RB69 highlights thermophilic adaptations, which include the presence of two disulfide bonds and an enhanced electrostatic complementarity at the DNA-protein interface. In contrast to gp43, several loops in the exonuclease and thumb domains are more closely packed; this apparently blocks primer binding to the exonuclease active site. A physiological role of this "closed" conformation is unknown but may represent a polymerase mode, in contrast to an editing mode with an open exonuclease site. This archaeal B DNA polymerase structure provides a starting point for structure-based design of polymerases or ligands with applications in biotechnology and the development of antiviral or anticancer agents.
Organizational Affiliation:
Abteilung Strukturforschung, Max-Planck-Institut für Biochemie, D-82152 Martinsried, Germany. hopfner@scripps.edu