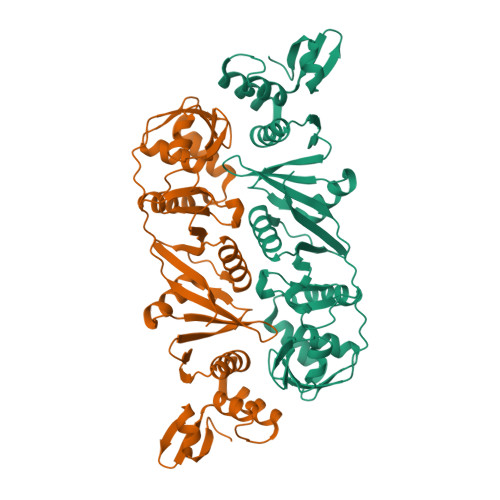
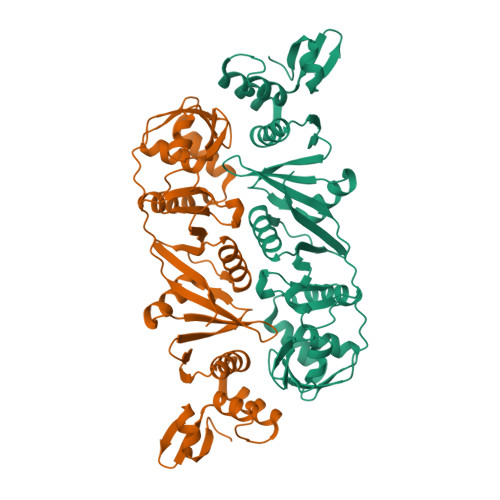
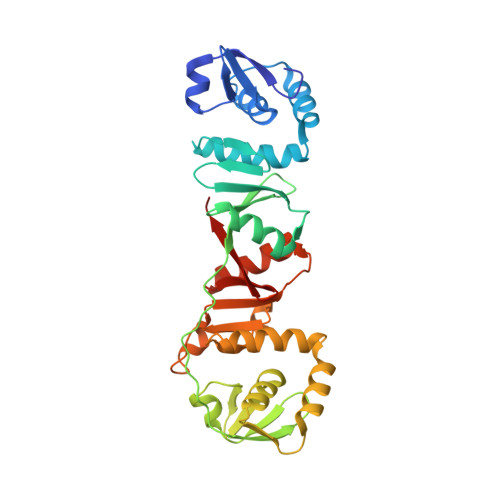
Crystal structure of a dimeric Archaeal Splicing Endonuclease
Li, H., Abelson, J.(2000) J Mol Biology 302: 639-648
- PubMed: 10986124
- DOI: https://doi.org/10.1006/jmbi.2000.3941
- Primary Citation of Related Structures:
1RLV - PubMed Abstract:
The splicing endonuclease from Archaeoglobus fulgidus (AF) belongs to the homodimeric family of splicing endonucleases, thought to have evolved from the homotetrameric endonucleases. We report here the crystal structure of the AF endonuclease determined at 2.8 A. The crystal structure of the full-length AF endonuclease contains a homodimer, with each monomer consisting of two homologous repeats joined together by an extended polypeptide chain of ten amino acid residues. The C-terminal repeat has a strong homology to that of a single subunit of the previously determined homotetrameric tRNA splicing endonuclease from Methanococcus jannaschii (MJ), indicating its role in catalysis. The N-terminal repeat is a more degenerate form of the MJ enzyme. Thus the N-terminal repeat is a "non-active" endonuclease fold evolved from the "active" one. By detailed comparison of the structures of the N-terminal and the C-terminal repeats, the binding region for RNA substrates containing a bulge-helix-bulge motif can be identified. Based on the identified RNA-binding region, a cation-pi interaction is suggested to be responsible for coordinating activities between the two active sites. In addition, the full-length AF endonuclease can adopt a higher-ordered fibrous structure in solution, as revealed by the unusual crystallographic packing interactions and other biochemical analysis. This 4(3)-fold fibrous structure adopted by the full-length enzyme is inaccessible to the RNA substrate and is largely stabilized by the first 60 amino acid residues. A mutated form of AF endonuclease with its first 60 residues removed catalyzes the cleavage reaction at a significantly higher rate. Whether there is any role in vivo for this structure-mediated modulation of activity remains to be determined.
Organizational Affiliation:
Division of Biology, California Institute of Technology, Pasadena, CA, USA. hongli@sb.fsu.edu