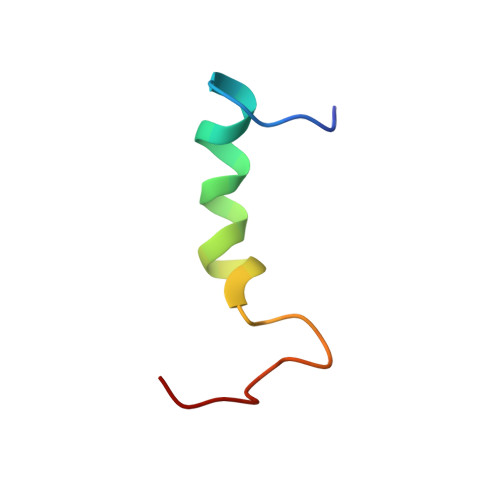
NMR solution structure of a peptide from the mdm-2 binding domain of the p53 protein that is selectively cytotoxic to cancer cells
Rosal, R., Pincus, M.R., Brandt-Rauf, P.W., Fine, R.L., Michl, J., Wang, H.(2004) Biochemistry 43: 1854-1861
- PubMed: 14967026
- DOI: https://doi.org/10.1021/bi035718g
- Primary Citation of Related Structures:
1Q2F, 1Q2I - PubMed Abstract:
We have recently found that a peptide from the mdm-2 binding domain of the p53 protein induced rapid membranolytic necrosis of a variety of different human cancer cell lines. To determine the role of solution structure in this peptide's selective and rapid tumor membrane disruptive behavior, we have performed two-dimensional NMR on a 32-residue sequence called PNC-27, in both an aqueous cytosolic-like and a mixed organic membrane-mimetic solution environment. In an aqueous milieu, PNC-27 contains three alpha-helical domains connected by loop structures, forming an S shape, and another similar structure with less helical structure. In a solution environment simulating a membrane, the helical domains found in water increase in length, forming three classes of structures, all of which form a U-shaped helix-coil-helix ensemble. In both solvent systems, this peptide forms amphipathic structures such that its hydrophobic residues coalesce on one face while the polar residues aggregate on the opposite face. The ability to form these unique structures in these two solution environments may allow the PNC-27 peptide to selectively and rapidly disrupt cancer cell membranes.
Organizational Affiliation:
Department of Environmental Health Sciences, Mailman School of Public Health of Columbia University, 60 Haven Avenue, New York, New York 10032, USA.